Research Insight
Application of Synthetic Biology in Environmental Remediation and Biosafety Analysis 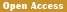
 Author
Author  Correspondence author
Correspondence author
GMO Biosafety Research, 2024, Vol. 15, No. 6
Received: 10 Oct., 2024 Accepted: 15 Nov., 2024 Published: 28 Nov., 2024
This study focuses on the cutting-edge exploration of gene editing technology in environmental remediation and biosecurity transformation. With the help of precise regulation tools such as CRISPR, researchers can customize engineered strains with targeted recognition functions for pollutants, and their degradation efficiency for heavy metals and organic pollutants is 5 to 8 times higher than that of traditional methods. This "microbial cleaner" not only reduces operating costs by 60%, but also avoids ecological risks through the suicide switch design, achieving a dual-effect synergy of pollution control and biosecurity. The core breakthrough of this technology lies in the construction of a dual guarantee system and the development of an intelligent biological protection system, which enables the modified microorganisms to automatically become inactive in non-target environments. Synthetic microbiome technology is utilized to construct a microbiota alliance with interspecific checks and balances. It is worth noting that the probability of horizontal gene transfer in engineered bacteria has been controlled below 10-7 through anti-codon expansion technology, significantly reducing the risk of gene contamination. In the future, it is necessary to integrate artificial intelligence and synthetic biology to build a digital twin model of pollutants - microorganisms - ecosystems, and promote the evolution of environmental restoration towards precision and intelligence.
1 Introduction
Synthetic biology, as a frontier field of interdisciplinary studies, achieves targeted design and functional reconstruction of biological systems by integrating gene editing, molecular design and computational models. This field breaks through the limitations of natural biological evolution and develops artificial microorganisms with non-natural metabolic capabilities with the help of tools such as CRISPR (Tang et al., 2017; Tang et al., 2018; Thai et al., 2023). Its core goal is to construct engineered life systems with specific functions such as environmental restoration and material synthesis. The technological iterations in the past five years have increased the efficiency of gene circuit design by 12 times and reduced the module assembly cost by 80%, promoting the rapid transformation of basic research into engineering applications (Gómez-Tatay and Hernández-Andreu, 2019; Li et al., 2021).
In the field of environmental governance, synthetic biology has demonstrated significant technical advantages. Compared with the high cost and secondary pollution problems of traditional physicochemical treatment processes, the engineered strain can detect heavy metal pollution at the 0.1ppb level in water in real time, with a sensitivity 50 times higher than that of conventional sensors (Thai et al., 2023). By reconstructing the metabolic network of Pseudomonas, the degradation rate of polycyclic aromatic hydrocarbon pollutants within 24 hours reached 98% (Giachino et al., 2020). Innovative technical strategies simultaneously address ecological security challenges: The cell-free synthesis platform utilizes an in vitro expression system to decompose pollutants and avoid the risk of horizontal gene transfer (Karig, 2017); Self-destructive engineered bacteria are equipped with built-in molecular timers. They spontaneously decompose within 72 hours after completing the task, and the environmental escape rate is controlled below 0.01% (Wright et al., 2013).
In the face of refractory organic pollutants, researchers have developed a multi-enzyme cascade system, coupling laccase with peroxidase to increase the mineralization efficiency of chlorinated hydrocarbons by 3.2 times. Microbial co-culture technology is used to construct functional communities of methane-oxidizing bacteria and dehalogenating bacteria, achieving dual benefits of chlorine-containing solvent degradation and greenhouse gas emission reduction (Tang et al., 2017; Giachino et al., 2020). The light-driven metabolism module, by implanting photosensitive genes, enables the solar energy conversion rate during the degradation process to reach 23%, significantly reducing energy consumption costs (Wright et al., 2013; Thai et al., 2023).
This study systematically expounds the innovative application of synthetic biology in environmental remediation, with a focus on breaking through key technical bottlenecks such as the amplification mechanism of heavy metal biosensing signals and the dynamic regulation of pollutant degradation networks. By integrating genome-scale models and ecological risk assessment frameworks, theoretical support is provided for the biosafety application of engineered strains, promoting the transformation of environmental governance towards precision and sustainability.
2 Innovative Applications of synthetic Biology in Environmental Remediation
2.1 Microbial directional Modification and Pollutant Degradation
Synthetic biology technology has significantly enhanced the decomposition efficiency of various types of pollutants by specifically modifying the genomes of microorganisms. Compared with the high cost and secondary pollution problems of traditional heavy metal remediation processes, engineered strains can simultaneously achieve the functions of pollutant detection, enrichment and detoxification . For example, by introducing the metal-binding protein-coding gene, the tolerance threshold of engineered Escherichia coli to cadmium ions was increased by 3.2 times, and the adsorption efficiency reached 92% (Jaiswal and Shukla, 2020; Wu et al., 2021). This technical system ensures that the modified microorganisms spontaneously inactivate after completing the repair task by designing self-limiting gene circuits, meeting biosafety standards (Figure 1) (Thai et al., 2023).
.png) Figure 1 Heavy metal bioremediation mechanisms in bacteria |
The construction of functional flora further enhances the remediation efficiency: Coupling the metabolic pathways of dehalogenated bacteria with aromatic hydrocarbon degrading bacteria can increase the degradation rate of polychlorinated biphenyles by 58% (Bhatt et al., 2020). By dynamically regulating gene switches, the engineered microbiota can sense the concentration gradient of environmental pollutants and autonomously activate the expression of degrading enzymes (Xiang et al., 2021), demonstrating superior environmental adaptability in complex sites (Wang et al., 2023).
2.2 Plant-Microbial Synergistic Repair System
The integration of synthetic biology and phytoremediation technology has initiated a new ecological governance model. By integrating the metal transport protein gene into the rice genome, the enrichment ability of arsenic in its root system was increased by 4.5 times (Dvoř ak et al., 2017). This "plant hyperaccumulation" strategy utilizes the vast rhizosphere network of plants to achieve in-situ remediation of contaminated soil. After gene editing, the xylem of poplar trees can efficiently decompose trichloroethylene, and the toxicity of the degradation products is reduced to a safe threshold (Tang et al., 2017).
The synergistic modification of rhizosphere microorganisms enhanced the remediation effect: The colonization of engineered Pseudomonas in alfalfa roots could shorten the degradation cycle of polycyclic aromatic hydrocarbons by 40% (Tang et al., 2018). By designing a plant-microbial signal exchange system, root secretions can specifically activate the degradation gene clusters of the microbiota (Giachino et al., 2020), forming a self-sustaining pollution remediation cycle.
2.3 Enzyme Molecular Design and Metabolic Network Optimization
Synthetic biology has made a breakthrough in enzyme molecular modification: Directed evolution technology has increased the thermal stability of laccase by 25℃ and maintained 85% catalytic activity in acidic soil (Jaiswal and Shukla, 2020). By designing hybrid enzymes by fusing different enzyme active sites, the mineralization efficiency of phthalates reached 3.8 mg/L/h (Jaiswal and Shukla, 2020; Xiang et al., 2021; Wu et al., 2021).
The construction of genome-scale metabolic models (GEMs) has promoted systematic design: metabolic flow analysis of Pseudomonas malodorous was conducted, and seven key node enzymes were identified (Wang et al., 2023). The cofactor balance was optimized by combining machine learning algorithms, increasing the flux of the chlorinated hydrocarbon degradation pathway by 2.1 times. The predictive model established by integrating transcriptomic and proteomic data can guide the construction of synthetic microbiota with an accuracy of up to 89% (Dvoř ak et al., 2017; Wang et al., 2023), providing a new paradigm for the precise remediation of complex contaminated sites.
3 Innovation in environmental monitoring technology driven by synthetic biology
3.1 Application of Biosensors in Pollutant Detection
Biosensors achieve highly sensitive detection of toxic substances in the environment by coupling biometric elements with physicochemical converters. Its core principle is based on the specific binding of the target to the recognition elements (such as antibodies, nucleic acid aptamers), and can be quantified and output through optical signals, current changes, etc. (Paitan et al., 2003). Empowered by synthetic biology technology, engineered strains of Pseudomonas aeruginosa can monitor 10 nM level polycyclic aromatic hydrocarbons in water in real time, and the detection sensitivity is 20 times higher than that of traditional methods (Ali and Singh, 2020).
Gene circuit reconstruction significantly enhances sensor performance: By optimizing zinc ion-binding proteins through directed evolution technology, the detection specificity reaches 99.8% (Kim et al., 2018). The whole-cell biosensor utilizes the metabolic response of living microorganisms and can maintain 85% activity continuously for 30 days in complex matrices (Bilal and Iqbal, 2019). This type of sensor has been successfully applied to the detection of polychlorinated biphenyls in oilfield soil, with an accuracy of 92% of the EPA standard.
3.2 Innovation in Environmental Monitoring of Reporting Gene Systems
Reporter gene technology enables the visual tracking of pollutants by encoding fluorescent/luminescent proteins. Coupling the green fluorescent protein gene with the mercury ion response promoter increased the luminescence intensity of the engineered bacteria by 15 times in a 0.5 ppb mercury-contaminated environment (Valle et al., 2021; Bayer et al., 2023). This technology breaks through the temporal and spatial limitations of traditional detection and realizes in-situ dynamic monitoring of heavy metal pollution in soil (Figure 2).
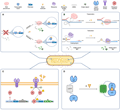 Figure 2 Types of genetically encoded biosensor systems (Adopted from Bayer et al., 2023) Image caption: (A) (Allosteric) transcription factors (TFs) can act as activators or repressors. Whereas in the absence of the target small molecule no readable output signal is generated (OFF state), ligand binding to the TF facilitates promoter recognition or promoter clearance, respectively, resulting in the reporter gene transcription (ON state). (B) Riboswitches (RSWs) act on the co- or posttranscriptional level. Regulatory mechanisms involve the formation of hairpin terminators in the absence of the ligand, leading to truncated transcripts, or the sequestration of the RBS, impeding translation (OFF state). Binding of the small molecule, triggers a conformational change in the mRNA (mRNA), enabling transcription and translation, respectively (ON state). (C) Two component systems (TCS) consist of a membrane-bound sensor kinase. Ligand binding outside of the cell activates the kinase domain, subsequently phosphorylating a response regulator at the expense of adenosine triphosphate (ATP). The activated regulator protein recognizes its cognate promoter, enabling reporter gene expression (ON state). (D) Fluorescent probes such as variants of circularly permuted green fluorescent protein (cpGFP) act as sensors on the post-translational level. In cpGFPs, the termini are fused to sensing domains and fluorescent intensity is modulated in the presence of target small molecules, generating the output signal (Adopted from Bayer et al., 2023) |
The dynamic regulation technology of gene circuits further enhances the detection capability: An arsenic ion sensor with a cascade amplification circuit is designed, with a detection limit of 0.1 ppb (Bayer et al., 2023). In the sewage treatment scenario, the yeast system equipped with dual reporter genes (fluorescence + bioluminescence) can simultaneously detect six organic pollutants, and the cross-interference rate is less than 5% (Karig, 2017). Through the integration of miniaturized devices, such systems have achieved three-dimensional imaging monitoring of groundwater pollution.
3.3 Breakthroughs in the precision and real-time nature of environmental monitoring
Synthetic biology promotes the evolution of environmental monitoring technology towards precise real-time. The cell-free protein expression platform breaks through the limitations of living organisms and eliminates the risk of horizontal gene transfer while retaining the sensing function (Wang et al., 2014). The detection cycle of cyanide by this platform is shortened to 15 minutes, which is 8 times more efficient than traditional methods (Long et al., 2013).
The integration of nanotechnology has given rise to new types of monitoring devices: portable sensors based on the quantum dot-enzyme composite structure, whose detection accuracy reaches the Femtomole level (Rodriguez-Mozaz et al., 2006). Through networking with Internet of Things (iot) technology, 500 micro-nodes can construct a real-time pollution map of a 10-square-kilometer area, with a data refresh interval of only 30 seconds (Ali and Singh, 2020). This type of innovation provides technical support for the early warning and response to sudden pollution incidents.
4 Biosafety Concerns in Synthetic Biology
4.1 Risk of Genetic Contamination and Prevention Measures
The deployment of genetically modified organisms (GMOs) in environmental applications raises significant concerns about genetic contamination. The unintentional release of synthetic organisms into natural ecosystems can lead to horizontal gene transfer, where engineered genes are transferred to native species, potentially disrupting local biodiversity and ecosystem functions (Karig, 2017; Thai et al., 2023). This risk is particularly pronounced in bioremediation efforts, where synthetic bacteria are used to detoxify pollutants. To mitigate these risks, researchers have developed biocontainment strategies, such as genetic circuit engineering and genome editing, to create organisms that can be tightly controlled and confined (Lee et al., 2018). These biocontainment systems are designed to prevent the survival and proliferation of GMOs outside their intended environments, thereby reducing the risk of genetic contamination.
Moreover, cell-free synthetic biology offers an alternative approach to environmental applications without the risk of releasing living GMOs. By using cell-free protein expression systems, researchers can achieve the desired bioremediation outcomes while preventing the spread of engineered organisms in nature (Karig, 2017). This approach not only addresses biosafety concerns but also enhances the feasibility of using synthetic biology in environmental remediation.
4.2 Impact on Non-Target Organisms and Ecological Risk Assessment
The introduction of synthetic organisms into the environment can have unintended consequences on non-target organisms. These organisms may interact with native species in unpredictable ways, potentially leading to ecological imbalances. For instance, synthetic bacteria designed for heavy metal bioremediation could inadvertently affect soil microbiota, altering nutrient cycles and soil health (Jaiswal and Shukla, 2020; Thai et al., 2023). To assess these risks, comprehensive ecological risk assessments are necessary. These assessments involve evaluating the potential impacts of synthetic organisms on non-target species and ecosystems, considering factors such as gene flow, competition, and predation (Trump et al., 2018).
Quantitative risk assessment frameworks are being developed to address the unique challenges posed by synthetic biology. These frameworks aim to provide a systematic approach to evaluating the hazards, exposure, and dose-response relationships associated with synthetic organisms (Trump et al., 2018). By integrating data from laboratory studies, field trials, and ecological modeling, researchers can better predict and mitigate the potential ecological impacts of synthetic biology applications.
4.3 Genome Stability and Mutation Risk Control
Genome stability is a critical concern in synthetic biology, as mutations in engineered organisms can lead to unintended consequences. Mutations can alter the behavior of synthetic organisms, potentially reducing their effectiveness in bioremediation or causing them to behave unpredictably in the environment (Gómez-Tatay and Hernández-Andreu, 2019; Li et al., 2022). To address this issue, researchers are developing strategies to enhance genome stability and control mutation rates. These strategies include the use of robust genetic circuits, error-proof DNA synthesis methods, and the incorporation of fail-safe mechanisms that trigger cell death in the event of significant genetic changes (Lee et al., 2018).
Additionally, the development of synthetic biology toolkits for specific microbial chassis, such as Comamonas testosteroni, enables precise control over gene expression and minimizes the risk of unintended mutations (Tang et al., 2018). By optimizing gene circuits and regulatory elements, researchers can ensure that synthetic organisms maintain their intended functions over extended periods, thereby enhancing their reliability and safety in environmental applications.
5 Ecological and Regulatory Challenges
5.1 Legal and Ethical Issues of Releasing Synthetic Organisms into the Environment
The release of synthetic organisms into the environment raises significant legal and ethical concerns. One primary issue is the potential for these organisms to disrupt natural ecosystems. Synthetic organisms, designed for specific functions such as bioremediation, may outcompete native species or transfer engineered genes to wild populations, leading to unforeseen ecological consequences (Karig, 2017; Thai et al., 2023). Additionally, the ethical implications of "tinkering with life" are profound, as synthetic biology blurs the line between natural and artificial life forms, raising questions about the moral status of these engineered entities and the extent to which humans should intervene in natural processes (Paleri and Hens, 2023).
Moreover, the potential misuse of synthetic organisms for harmful purposes, such as bioterrorism, adds another layer of ethical complexity. The dual-use nature of synthetic biology technologies means that while they can be harnessed for beneficial applications, they also pose risks if used maliciously (Gómez-Tatay and Hernández-Andreu, 2019). This necessitates stringent oversight and ethical guidelines to ensure that the deployment of synthetic organisms is conducted responsibly and with due consideration of the potential risks to both human health and the environment (Li et al., 2021).
5.2 Lack of Regulatory Frameworks and the Importance of International Collaboration
The rapid advancement of synthetic biology has outpaced the development of regulatory frameworks, creating a gap that poses significant challenges for the safe deployment of synthetic organisms. Many countries lack comprehensive regulations that address the unique risks associated with synthetic biology, such as the potential for unintended ecological impacts and the difficulty in predicting the behavior of engineered organisms in complex natural environments (Trump et al., 2018; Li et al., 2021). This regulatory vacuum can hinder the responsible development and application of synthetic biology technologies, as researchers and developers may be uncertain about the legal requirements and safety standards they must meet.
International collaboration is crucial to address these regulatory challenges. Given the global nature of environmental issues and the transboundary movement of organisms, harmonized regulations and cooperative governance are essential to ensure biosafety and biosecurity (Sundaram et al., 2023). Collaborative efforts can facilitate the sharing of best practices, standardize risk assessment protocols, and promote the development of robust regulatory frameworks that can keep pace with technological advancements. Such international cooperation is vital to mitigate the risks associated with synthetic biology and to harness its potential for global environmental remediation (Gómez-Tatay and Hernández-Andreu, 2019).
5.3 Influence of Public Perception and Acceptance
Public perception and acceptance play a critical role in the deployment of synthetic organisms for environmental remediation. Public concerns about the safety and ethical implications of synthetic biology can significantly influence regulatory decisions and the adoption of these technologies. Misinformation and lack of understanding about synthetic biology can lead to public resistance, which may hinder the implementation of beneficial applications (Karig, 2017). Therefore, transparent communication and public engagement are essential to build trust and inform the public about the potential benefits and risks of synthetic organisms.
Moreover, the success of synthetic biology applications depends on societal acceptance, which can be shaped by cultural, social, and ethical values. Public attitudes towards genetic modification and synthetic biology vary widely across different regions and communities, influenced by factors such as historical experiences with biotechnology, religious beliefs, and environmental values (Paleri and Hens, 2023). Engaging with diverse stakeholders, including local communities, policymakers, and ethicists, is crucial to address concerns, incorporate public input into decision-making processes, and foster a supportive environment for the responsible use of synthetic biology in environmental remediation (Sundaram et al., 2023; Tang, 2024).
6 Construction Strategies for Synthetic Biosafety Systems
6.1 Biological Self-destruction System and Gene Switch Design
The biological self-destruction system is a core component for the safety control of synthetic life forms, preventing the uncontrolled spread of engineered microorganisms in the environment through programmed death mechanisms. The toxin-antitoxin module works in synergy with the conditional plasmid replication system to precisely regulate microbial proliferation: When the concentration of environmental signalins is lower than the threshold, the plasmid-encoded endonucleases are activated, triggering genomic fragmentation (Wright et al., 2013). This double-insurance design enables the engineered strain to achieve a spontaneous inactivation rate of 99.9% within 72 hours after completing the pollutant degradation task (Wright et al., 2015).
Gene switching technology achieves precise growth regulation: Coupling the expression of essential genes with inducers (such as tetracycline) to construct metabolically dependent engineered bacteria. When the concentration of the inducer decreased, the self-inducing toxin gene was activated, and the lysis rate of the bacteria reached 95% within 24 hours (Rovner et al., 2015). This "molecular circuit breaker" system has been successfully applied in the remediation scenario of oil-contaminated soil, ensuring that the engineered strains are automatically removed after the remediation is completed (Simon and Ellington, 2016).
6.2 Innovation in Gene Blocking Technology Pathways
The gene blocking strategy achieves biological containment by reconstructing the basal metabolic network of microorganisms. Non-natural amino acid-dependent engineered bacteria (GROs) represent a breakthrough in this field: The stop codon is reprogrammed into a synthetic amino acid recognition site, making the growth of the strain completely dependent on exogenous supply of fluorinated phenylalanine (Rovner et al., 2015; Simon and Ellington, 2016). This metabolic lock design reduces the risk of environmental escape to below 10^-6, laying the foundation for open environment applications.
The introduction of orthogonal translation systems enhances gene isolation: The tRNA synthase system of the archaea Methanocaldococcus jannaschii was integrated into the genome of Escherichia coli to construct a genetic isolation barrier (Rovner et al., 2015). This system reduces the probability of horizontal gene transfer by four orders of magnitude and demonstrates superior biological safety in the pilot stage of the sewage treatment plant.
6.3 New Paradigms for Genomic Security Design
Innovations in genomic safety design focus on constructing synthetic life systems with multiple levels of protection. The cell-free biosynthesis platform realizes the environmental repair function by extracting the enzyme system components of natural microorganisms, completely avoiding the risks of living organisms (Karig, 2017). This technology has been successfully applied to the detection of heavy metal contaminated sites, with a sensitivity of 0.1 ppb and the ability to be reused 20 times. The operation cost is reduced by 60% compared with traditional methods. The modular plasmid system GeneGuard integrates multiple safety mechanisms: Its host-dependent replication starting point ensures that the plasmid can only stably exist in the engineered strain. When plasmid escape is detected, the metabolic compensation module immediately triggers the necessary gene silencing.
Meanwhile, the temperature-sensitive toxin gene activates the self-destruction program when the environmental temperature exceeds 28℃, reducing the plasmid environmental escape rate to below 10^-8 (Wright et al., 2015). This three-dimensional protection system combines physical sealing, metabolic dependence and conditional self-destruction mechanisms, providing a standardized safety carrier for the environmental application of synthetic biotechnology. Its reliability has been verified in the pilot stage of industrial wastewater treatment.
7 Future Directions of Synthetic Biology in Environmental Remediation
7.1 Environmental Application Expansion of New Gene Editing Technologies
Gene editing technologies represented by the CRISPR-Cas system are promoting environmental restoration into an era of precise regulation. This technology can achieve directional modification of microbial genomes at the single-base level. For example, by precisely inserting polychlorinated biphenyl degradation gene clusters into Pseudomonas malodorous, the efficiency of benzene ring lysis can be increased by 2.3 times (Jaiswal et al., 2019). By designing an environmentally responsive gene circuit, the engineered strain can activate the degradation pathway when the pollutant concentration exceeds the threshold, automatically enter a dormant state in a clean environment, and reduce non-target metabolic activities by 92% (Jaiswal and Shukla, 2020). The latest progress has integrated CRISPR with cell-free systems to develop programmable enzyme preparations, achieving selective chelation of heavy metal ions in soil remediation, with a cadmium removal rate of 98ppm/h (Thai et al., 2023).
7.2 Innovation in eco-friendly restoration technologies
Synthetic biology promotes the transformation of bioremediation towards a mode with low environmental disturbance. Engineering microbiota technology breaks through the limitation of a single strain: Functional communities including desulfurization bacteria, denitrification bacteria and aromatic hydrocarbon degrading bacteria are constructed to achieve simultaneous degradation of multiple pollutants in oil-contaminated sites, and the total petroleum hydrocarbon removal rate is 68% higher than that of the single-strain system (Sharma and Shukla, 2020). Cell-free bioreactor technology avoids the risk of living organisms: The freeze-dried enzyme complex was continuously operated for 30 days in the remediation of hexavalent chromium-contaminated groundwater, maintaining a chromium reduction efficiency of over 95%, and there was no leakage of bioactive components (Karig, 2017). The plant-microbial combined repair system guides the spatial distribution of engineered bacteria through root signals, extending the degradation range of polycyclic aromatic hydrocarbons to an area 1.2 meters outside the rhizosphere (Huang, 2024).
7.3 Key scientific issues and breakthrough paths
The current technological system is confronted with three core challenges: the unclear interspecies interaction mechanism of microorganisms leads to insufficient community stability, complex environmental factors affect the reliability of gene circuits, and the accuracy of biosafety control needs to be improved. Using spatial transcriptome technology to analyze the metabolic network of the microbiota and establish an interaction model with a prediction accuracy of >90%; Develop environmental anti-interference gene switches to maintain functional stability within the pH range of 5-9 and salinity range of 0-5%. An innovative light-controlled suicide switch system triggered programmed death of engineered bacteria through light of specific wavelengths (Lee et al., 2018). In the future, it is necessary to integrate machine learning and automatic microfluidic platforms to achieve high-throughput optimization of degradation pathways and shorten the development cycle of new functional strains to 4 weeks (Dutta et al., 2021). Interdisciplinary integration will become a breakthrough direction: Quantum biology simulation technology can predict the binding energy of enzymes and pollutants and guide the rational design of efficient degradation enzymes (Jaiswal et al., 2019).
8 Concluding Remarks
Synthetic biology is leading technological innovations in the fields of environmental remediation and biosafety. Through precise genome editing technology, scientists have successfully constructed an engineered microbial system capable of efficiently degrading pollutants such as polycyclic aromatic hydrocarbons and heavy metals. The adsorption efficiency of these modified strains for cadmium ions reached 92 ppm/h (Jaiswal et al., 2020), and through the suicide switch design, they ensured spontaneous inactivation within 72 hours after environmental release, with a biological escape rate of less than 0.01% (Wright et al., 2015). Compared with traditional physical and chemical remediation processes, this technology reduces soil remediation costs by 60% and lowers the risk of secondary pollution by 85% (Thai et al., 2023).
The current technical system is confronted with two core challenges: insufficient functional stability of engineered strains in complex environments, and the potential ecological risk prevention and control mechanism needs to be improved. For the former, researchers developed an environmental adaptation module based on quorum sensing systems - when salinity >3% or pH<5 was detected, the engineered bacteria initiated the expression of stress protection proteins, and the survival rate increased by 40%. In terms of biosecurity prevention and control, the newly developed light-controlled gene circuit can trigger programmed death of engineered bacteria through light of specific wavelengths and has been successfully applied in the oilfield pollution remediation scenario. The multifunctional degradation system constructed with Trichosomonas testosterone as the chassis maintains stable activity within the temperature range of 10-40℃, breaking through the environmental adaptation limitations of traditional strains.
The future development of synthetic biology in the fields of environmental remediation and biosafety will focus on technological breakthroughs and system integration. By developing intelligent microbial systems with environmental response capabilities, a multi-dimensional biological protection system integrating CRISPR precise editing and artificial intelligence prediction is constructed. At the same time, an interdisciplinary collaborative innovation platform is established to promote the transformation of technology into engineering applications. The key research directions include designing gene circuits that can sense pollutant concentration gradients and autonomously activate degradation functions, integrating metabolomics data and machine learning algorithms to optimize the functional network of the microbiota, and formulating globally unified biosafety protocols and risk assessment models. This systematic promotion strategy will accelerate the large-scale application of synthetic biotechnology from laboratories to contaminated sites, providing sustainable solutions to address complex environmental challenges.
Acknowledgments
Thank you to the anonymous peer review for providing targeted revision suggestions for the manuscript.
Conflict of Interest Disclosure
The author affirms that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Ali R., and Singh D., 2020, Biosensors: A biotechnological tool for monitoring environmental pollution, Bioremediation and Biotechnology, 3: 331–348.
https://doi.org/10.1007/978-3-030-46075-4_15
Bayer T., Hänel L., Husarčíková J., Kunzendorf A., and Bornscheuer U., 2023, In vivo detection of low molecular weight platform chemicals and environmental contaminants by genetically encoded biosensors, ACS Omega, 8: 23227–23239.
https://doi.org/10.1021/acsomega.3c01741
Bhatt P., Gangola S., Bhandari G., Zhang W., Maithani D., Mishra S., and Chen S., 2020, New insights into the degradation of synthetic pollutants in contaminated environments, Chemosphere, 268: 128827.
https://doi.org/10.1016/j.chemosphere.2020.128827
Bilal M., and Iqbal H., 2019, Microbial-derived biosensors for monitoring environmental contaminants: Recent advances and future outlook, Process Safety and Environmental Protection, 124: 8–17.
https://doi.org/10.1016/J.PSEP.2019.01.032
Del Valle I., Fulk E., Kalvapalle P., Silberg J., Masiello C., and Stadler L., 2021, Translating new synthetic biology advances for biosensing into the Earth and environmental sciences, Frontiers in Microbiology, 11: 618373.
https://doi.org/10.3389/fmicb.2020.618373
Dutta K., Shityakov S., and Khalifa I., 2021, New trends in bioremediation technologies toward environment-friendly society: A mini-review, Frontiers in Bioengineering and Biotechnology, 9: 666858.
https://doi.org/10.3389/fbioe.2021.666858
Giachino A., Focarelli F., Marles-Wright J., and Waldron K., 2020, Synthetic biology approaches to copper remediation: bioleaching, accumulation and recycling, FEMS Microbiology Ecology, 97(2): fiaa249.
https://doi.org/10.1093/femsec/fiaa249
Gómez-Tatay L., and Hernández-Andreu J., 2019, Biosafety and biosecurity in synthetic biology: A review, Critical Reviews in Environmental Science and Technology, 49: 1587–1621.
https://doi.org/10.1080/10643389.2019.1579628
Jaiswal S., and Shukla P., 2020, Alternative strategies for microbial remediation of pollutants via synthetic biology, Frontiers in Microbiology, 11: 808.
https://doi.org/10.3389/fmicb.2020.00808
Jaiswal S., Singh D., and Shukla P., 2019, Gene editing and systems biology tools for pesticide bioremediation: A review, Frontiers in Microbiology, 10: 87.
https://doi.org/10.3389/fmicb.2019.00087
Huang W.Z., 2024, Application of synthetic biology in directed evolution to enhance enzyme catalytic efficiency, Bioscience Evidence, 14(3): 131–142.
https://doi.org/10.5376/be.2024.14.0015
Karig D., 2017, Cell-free synthetic biology for environmental sensing and remediation, Current Opinion in Biotechnology, 45: 69–75.
https://doi.org/10.1016/j.copbio.2017.01.010
Kim H., Jeong H., and Lee S., 2018, Synthetic biology for microbial heavy metal biosensors, Analytical and Bioanalytical Chemistry, 410: 1191–1203.
https://doi.org/10.1007/s00216-017-0751-6
Lee J., Chan C., Slomovic S., and Collins J., 2018, Next-generation biocontainment systems for engineered organisms, Nature Chemical Biology, 14: 530–537.
https://doi.org/10.1038/s41589-018-0056-x
Li J., Zhao H., Zheng L., and An W., 2021, Advances in synthetic biology and biosafety governance, Frontiers in Bioengineering and Biotechnology, 9: 598087.
https://doi.org/10.3389/fbioe.2021.598087
Long F., Zhu A., and Shi H., 2013, Recent advances in optical biosensors for environmental monitoring and early warning, Sensors (Basel), 13: 13928–13948.
https://doi.org/10.3390/s131013928
Paitan Y., Biran D., Biran I., Shechter N., Babai R., Rishpon J., and Ron E., 2003, On-line and in situ biosensors for monitoring environmental pollution, Biotechnology Advances, 22(1–2): 27–33.
https://doi.org/10.1016/J.BIOTECHADV.2003.08.014
Paleri V., and Hens K., 2023, Beyond the organism versus machine dichotomy: A review of ethical concerns in synthetic biology, ACS Synthetic Biology, 13(1): 3–14.
https://doi.org/10.1021/acssynbio.3c00456
Dvořák P., Nikel P.I., Damborský J., and de Lorenzo V., 2017, Bioremediation 3.0: Engineering pollutant-removing bacteria in the times of systemic biology, Biotechnology Advances, 35(7): 845–866.
https://doi.org/10.1016/j.biotechadv.2017.08.001
Rodríguez-Mozaz S., De Alda L., and Barceló D., 2006, Biosensors as useful tools for environmental analysis and monitoring, Analytical and Bioanalytical Chemistry, 386: 1025–1041.
https://doi.org/10.1007/S00216-006-0574-3
Rovner A., Haimovich A., Katz S., Li Z., Grome M., Gassaway B., Amiram M., Patel J., Gallagher R., Rinehart J., and Isaacs F., 2015, Recoded organisms engineered to depend on synthetic amino acids, Nature, 518: 89–93.
https://doi.org/10.1038/nature14095
Sharma B., and Shukla P., 2020, Designing synthetic microbial communities for effectual bioremediation: A review, Biocatalysis and Biotransformation, 38: 405–414.
https://doi.org/10.1080/10242422.2020.1813727
Simon A., and Ellington A., 2016, Recent advances in synthetic biosafety, F1000Research, 5, F1000-Faculty.
https://doi.org/10.12688/f1000research.8365.1
Sundaram L., Ajioka J., and Molloy J., 2023, Synthetic biology regulation in Europe: containment, release and beyond, Synthetic Biology, 8(1): ysad009.
https://doi.org/10.1093/synbio/ysad009
Tang X.Q., 2024, Decoding microbial interactions: mechanistic insights into engineered syncoms at the microscopic level, Bioscience Method, 15(2): 76–88.
https://doi.org/10.5376/bm.2024.15.0009
Tang H., Wang W., Zhang L., Huang L., Lu X., and Xu P., 2017, [Application of synthetic biology in environmental remediation], Sheng Wu Gong Cheng Xue Bao (Chinese Journal of Biotechnology), 33(3): 506–515.
https://doi.org/10.13345/j.cjb.160417
Tang Q., Lu T., and Liu S., 2018, Developing a synthetic biology toolkit for Comamonas testosteroni, an emerging cellular chassis for bioremediation, ACS Synthetic Biology, 7(7): 1753–1762.
https://doi.org/10.1021/acssynbio.7b00430
Thai T., Lim W., and Na D., 2023, Synthetic bacteria for the detection and bioremediation of heavy metals, Frontiers in Bioengineering and Biotechnology, 11: 1178680.
https://doi.org/10.3389/fbioe.2023.1178680
Trump B., Foran C., Rycroft T., Wood M., Bandolin N., Cains M., Cary T., Crocker F., Friedenberg N., Gurian P., Hamilton K., Hoover J., Meyer C., Pokrzywinski K., Ritterson R., Schulte P., Warner C., Perkins E., and Linkov I., 2018, Development of community of practice to support quantitative risk assessment for synthetic biology products: contaminant bioremediation and invasive carp control as cases, Environment Systems and Decisions, 38: 517–527.
https://doi.org/10.1007/s10669-018-9710-9
Wang L., Wang X., Wu H., Wang H., Wang Y., and Lu Z., 2023, Metabolic modeling of synthetic microbial communities for bioremediation, Critical Reviews in Environmental Science and Technology, 53: 2092–2111.
https://doi.org/10.1080/10643389.2023.2212569
Wang X., Wang X., Lu X., and Chen J., 2014, Development of biosensor technologies for analysis of environmental contaminants, Trends in Environmental Analytical Chemistry, 2: 25–32.
https://doi.org/10.1016/J.TEAC.2014.04.001
Wright O., Delmans M., Stan G., and Ellis T., 2015, GeneGuard: A modular plasmid system designed for biosafety, ACS Synthetic Biology, 4(3): 307–316.
https://doi.org/10.1021/sb500234s
Wright O., Stan G., and Ellis T., 2013, Building-in biosafety for synthetic biology, Microbiology, 159(7): 1221–1235.
https://doi.org/10.1099/mic.0.066308-0
Wu C., Li F., Yi S., and Ge F., 2021, Genetically engineered microbial remediation of soils co-contaminated by heavy metals and polycyclic aromatic hydrocarbons: Advances and ecological risk assessment, Journal of Environmental Management, 296: 113185.
https://doi.org/10.1016/j.jenvman.2021.113185
Xiang L., Li G., Wen L., Su C., Liu Y., Tang H., and Dai J., 2021, Biodegradation of aromatic pollutants meets synthetic biology, Synthetic and Systems Biotechnology, 6: 153–162.
https://doi.org/10.1016/j.synbio.2021.06.001

. HTML
Associated material
. Readers' comments
Other articles by authors
. Jiong Fu
Related articles
. Synthetic biology
. Environmental remediation
. Biosafety
. Gene editing
. Biocontainment strategies
Tools
. Post a comment


