 Author
Author  Correspondence author
Correspondence author
GMO Biosafety Research, 2024, Vol. 15, No. 5
Received: 10 Aug., 2024 Accepted: 22 Sep., 2024 Published: 09 Oct., 2024
This study reviews the development of wheat transformation technologies, tracing their progress from early explorations in tissue culture and gene delivery to the current optimization and application of mainstream methods such as Agrobacterium-mediated transformation, particle bombardment, and CRISPR-based gene editing. It summarizes key achievements in the fields of disease resistance, abiotic stress tolerance, and nutritional quality enhancement. The research highlights that wheat’s complex hexaploid genome and relatively low transformation efficiency have long limited advances in functional gene research and molecular breeding. Presently, wheat transformation still faces major challenges, including strong genotype dependency, insufficient transformation stability, and limited public acceptance. With the rapid advancement of modern biotechnology, significant breakthroughs have been made in wheat transformation. Future development is expected to focus on genotype-independent transformation systems, high-throughput automation, and precision gene editing, while also addressing societal acceptance and biosafety concerns. This study provides important references for advancing functional genomics, precision breeding, and industrial applications of wheat.
1 Introduction
Wheat, as one of the world’s three major staple crops, plays a pivotal role in sustaining global food security and nutrition. Its widespread cultivation and consumption make it essential for the diets of billions of people worldwide. However, the genetic complexity of wheat—stemming from its large, hexaploid genome—has long posed significant challenges for researchers. This complexity, coupled with historically low transformation efficiency, has restricted advances in functional gene research and limited the pace of molecular breeding for trait improvement (Li et al., 2012; Kim et al., 2021).
With the rapid development of biotechnology, wheat transformation technologies have experienced continuous breakthroughs. Innovations such as morphogenic gene-mediated transformation, optimized Agrobacterium-mediated protocols, and advanced biolistic methods have substantially improved transformation efficiencies and expanded the range of transformable wheat genotypes. These advances are enabling more precise gene function studies, the development of genome-edited lines, and the introduction of valuable agronomic traits, including disease resistance and enhanced nutritional quality (Hayta et al., 2021; Johnson et al., 2023).
This study will systematically review the developmental history of wheat genetic transformation technologies, analyze the current application status, explore future directions for overcoming existing bottlenecks, and envision the broad application prospects of these transformation tools in wheat genetic improvement. The aim of this study is to provide essential reference for advancing wheat transformation technologies into a new stage of high efficiency and precision.
2 Origins and Early Exploration of Wheat Genetic Transformation
2.1 Establishment of early tissue culture and regeneration systems
Early efforts in wheat tissue culture explored a range of explants, including immature embryos, mature embryos, stem segments, and endosperm. Immature embryos quickly emerged as the most responsive explant for initiating morphogenic cultures and achieving high regeneration rates, though mature embryos and other tissues were also investigated for their potential (Kumar et al., 2017). Genotype played a significant role in tissue culture response, with some lines, such as CB037B, showing superior regeneration ability across explant types (Yin et al., 2011; Liu et al., 2022). Anther and microspore cultures provided additional avenues for generating regenerants and studying differentiation processes.
The success of wheat transformation has depended heavily on optimizing callus induction and plant regeneration protocols. Key factors included the choice of basal media (e.g., MS, N6, Aa), the use of organic additives and antioxidants, and, critically, the balance and concentration of plant growth regulators such as 2,4-D, dicamba, zeatin, and cytokinins (Yin et al., 2011). For example, picloram and 2,4-D were found to be effective auxins for callus induction, while the addition of zeatin and copper sulfate enhanced shoot regeneration. Hormone ratios and media composition were tailored to specific genotypes and explant types, resulting in regeneration frequencies as high as 100% in optimized systems (Kumar et al., 2017). The development of genotype-independent protocols and the use of computational models, such as artificial neural networks, further improved reproducibility and efficiency.
2.2 Initial attempts at gene delivery
The introduction of particle bombardment (biolistics) marked a pivotal advance in wheat genetic transformation. This method enabled the direct delivery of DNA into plant cells, bypassing some of the limitations of Agrobacterium-mediated transformation in wheat. Immature embryos were typically used as target tissues, with gold or tungsten particles coated with plasmid DNA carrying selectable marker genes (e.g., bar). Early protocols established the importance of pre-culturing explants and optimizing selection regimes to recover transgenic plants efficiently. The biolistic approach became widely adopted due to its applicability across diverse wheat genotypes and its ability to produce stable, heritable transgene integration (Sparks and Doherty, 2020).
Initial efforts to apply Agrobacterium-mediated transformation to wheat faced significant obstacles. Wheat’s recalcitrance to Agrobacterium infection, combined with its complex genome and low regeneration capacity, resulted in very low transformation efficiencies compared to other cereals like rice and maize. Early protocols required careful optimization of explant type, with immature embryos proving most responsive, and the use of specific Agrobacterium strains and vectors. Factors such as embryo size, pre-culture duration, co-cultivation conditions, and the presence of vir gene inducers like acetosyringone were found to significantly influence T-DNA delivery and plant regeneration (Kumar et al., 2019). Despite these optimizations, transformation frequencies in early studies often remained below 5%, and were highly genotype-dependent. Additionally, the integration of transgenes was sometimes unstable, and the recovery of fertile, morphologically normal plants was inconsistent.
2.3 Technical bottlenecks
A major technical bottleneck in wheat transformation has been the consistently low efficiency of Agrobacterium-mediated protocols, with reported rates often ranging from 0.3% to 5% in many genotypes. Only a few cultivars, such as ‘Fielder’ and ‘Bobwhite,’ have shown higher transformation rates, highlighting a strong genotype dependency that limits the broader application of these methods (Richardson et al., 2014; Kumar et al., 2019; Hayta et al., 2021). Furthermore, transgene insertion can be unstable, with variable copy numbers and occasional rearrangements, complicating downstream breeding and trait stability.
Another persistent challenge is the high frequency of chimerism—where only some cells in a regenerated plant carry the transgene—leading to phenotypic inconsistency and complicating the identification of true transgenic lines (Wu et al., 2003). Regenerated plants may also exhibit reduced fitness, abnormal morphology, or poor fertility, particularly in the primary (T0) generation, although these issues often diminish in subsequent generations (Hensel et al., 2017). These technical hurdles have necessitated ongoing refinement of transformation protocols and the search for more robust, genotype-independent systems (Kumar et al., 2019; Hayta et al., 2021).
3 Mainstream Transformation Strategies and Integration with Emerging Technologies
3.1 Improvements and expanded applications of agrobacterium-mediated transformation
Significant progress has been made in Agrobacterium-mediated transformation of wheat through the development of hypervirulent Agrobacterium strains and the optimization of inoculation methods, such as immersion techniques and precise control of co-culture duration. These refinements have led to substantial increases in transformation efficiency. For example, the use of the spring wheat cultivar ‘Fielder’ as a model genotype, combined with a centrifugation pretreatment and optimized co-cultivation, has enabled transformation efficiencies up to 33%, and even higher (up to 77.5%) when combined with developmental gene fusions like GRF-GIF. These advances have also been successfully adapted to other genotypes, such as ‘Cadenza’ and ‘Kronos’, though with lower efficiencies, indicating ongoing genotype dependency but broader applicability (Hayta et al., 2021). Additional protocol optimizations—including embryo size selection, pre-culture duration, and the use of vir gene inducers—have further improved T-DNA delivery and plant regeneration (Hu et al., 2003; Wu et al., 2003). The result is a robust, reproducible system that supports a wide range of applications, from overexpression and RNA interference to CRISPR-based genome editing (Hu et al., 2003).
3.2 Optimization of particle bombardment and integration with efficient regeneration systems
Particle bombardment remains a widely used method for wheat transformation, especially for genotypes less amenable to Agrobacterium infection. Recent refinements in this technique include the optimization of particle size, voltage settings, and DNA construct design, all of which have contributed to improved transformation efficiencies and more consistent results. The integration of particle bombardment with highly embryogenic callus systems has further increased the rate of positive transformants, as demonstrated by transformation efficiencies of 4% in some wheat varieties. The use of immature embryos and well-established regeneration protocols has enabled the successful transformation of a broad range of wheat genotypes, from elite cultivars to landraces, with fewer limitations related to genotype dependency compared to Agrobacterium-mediated methods. Additionally, advances in tissue culture, such as the use of basal segment calli and optimized selection media, have supported the recovery of fertile, stable transgenic plants.
3.3 Successful applications of genome editing in wheat
The CRISPR/Cas9 and Cas12a systems have been successfully adapted for wheat, with specific target design rules to address the crop’s polyploid genome and minimize off-target effects (Smedley et al., 2021). Tools like WheatCRISPR facilitate the design of highly specific guide RNAs, improving editing precision and efficiency (Cram et al., 2019). Multiplex genome editing-simultaneously targeting multiple genes or gene copies-has been achieved using tandemly arrayed tRNA–gRNA units and other strategies, enabling complex trait modification such as drought tolerance and increased grain size (Wang et al., 2018; Li et al., 2020; Abdallah et al., 2022).
A major breakthrough has been the development of DNA-free genome editing methods. Delivery of CRISPR/Cas9 as ribonucleoprotein (RNP) complexes or in vitro transcripts (IVTs) via particle bombardment or transient expression allows for efficient, transgene-free editing, reducing regulatory concerns and off-target mutations. These approaches can generate homozygous, stable mutants in the T0 generation and are applicable to both bread and durum wheat (Liang et al., 2018). Agrobacterium-mediated delivery of CRISPR/Cas9 has also been optimized, producing heritable edits and enabling both gene knockouts and knock-ins (Zhang et al., 2019).
3.4 Standardization of transgene detection and expression analysis tools
Detection and analysis of transgenes and genome edits in wheat have become more robust and standardized. Common selectable marker genes such as Bar, Hpt, and GFP are widely used, and their removal is facilitated by site-specific recombination systems like CRE/loxP, allowing for the generation of marker-free plants (Zhang et al., 2019). Molecular validation methods—including PCR, qRT-PCR, Southern blotting, and next-generation sequencing—are routinely employed to confirm the presence, absence, or precise editing of target genes (Liang et al., 2018). Rapid detection protocols, such as PCR-RFLP and restriction enzyme assays, enable efficient screening of edited lines (Kamiya et al., 2020). Transcriptomic analyses further support the assessment of gene expression changes and off-target effects.
4 Applications and Representative Achievements in Wheat Breeding
4.1 Disease-resistant transgenic wheat
Significant progress has been made in developing wheat varieties resistant to major diseases such as stripe rust, leaf rust, and powdery mildew. Strategies include the introduction of resistance (R) genes, engineering of nucleotide-binding leucine-rich repeat (NLR) receptors, and manipulation of susceptibility (S) genes using both traditional and genome editing approaches (Waites et al., 2025). The pyramiding of multiple resistance genes, as seen in cultivars like Guinong 29, has resulted in durable resistance to powdery mildew (Pm2, Pm21), stripe rust (Yr26), and leaf rust (Lr1, Lr46), while maintaining high agronomic performance. Transgenic approaches have also utilized pathogenesis-related (PR) proteins, such as PR1, to enhance both biotic and abiotic stress tolerance, with evidence of improved antioxidant enzyme activity and reduced pathogen susceptibility. The use of rice chitinase genes and other defense-related genes via transformation has further expanded the arsenal against fungal pathogens (Fahmy et al., 2018). CRISPR-based genome editing is now enabling precise and rapid development of disease-resistant lines, overcoming some limitations of conventional breeding (Waites et al., 2025).
4.2 Abiotic stress resistance (drought, salinity, heat)
Transgenic wheat lines carrying stress-responsive genes such as DREB and LEA have demonstrated outstanding performance under abiotic stresses including drought, salinity, and heat. For instance, overexpression of DREB genes significantly enhanced freezing tolerance, yield, and the stability of protein and starch quality. Studies have shown that the TaASR1-D gene functions by enhancing antioxidant capacity and activating the abscisic acid (ABA) signaling pathway. The overexpression (OE) lines exhibited greater sensitivity to ABA treatment, with significantly suppressed shoot and root growth, along with increased levels of ABA and H₂O₂, indicating the gene’s regulatory potential in response to abiotic stresses such as drought and salinity (Figure 1). Moreover, TaASR1-D notably improved grain yield under salt stress, conferring broad-spectrum stress resistance. In addition, the transcription factor ERF1-V, introduced from Haynaldia villosa, not only enhanced resistance to powdery mildew but also improved tolerance to salt and drought without compromising key agronomic traits. Multiple field trials have validated the stress adaptability of these transgenic materials under real-world environmental conditions (Qiu et al., 2021).
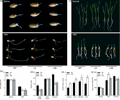 Figure 1 Overexpression of TaASR1-D enhanced ABA signalling (Adopted from Qiu et al., 2021) Image caption: (a) ABA sensitivity assay of primary root growth. (b) Root lengths of WT and OE seedlings under normal and ABA treatment conditions. (c) ABA sensitivity assay of seedling growth. (d) Relative values of shoot lengths, root lengths and biomasses after ABA treatment. (e) Relative ABA contents of WT and L39 under normal and stress treatment conditions. (f) H2O2 contents of WT and OE leaves under normal condition and H2O2, fluridone (Fd) + H2O2 treatment for 24 h. FW, fresh weight. All experiments were performed with at least three biological replicates. Data are means ± SE, and asterisks indicate significant differences between WT and OE lines using a t-test (*, P < 0.05; **, P < 0.01) (Adopted from Qiu et al., 2021) |
4.3 Nutritional and functional quality improvement
Efforts to enhance the nutritional profile of wheat have led to the development of high-protein, high-lysine, and iron-fortified transgenic lines. Mutational breeding and transgenic approaches have produced wheat with improved grain protein content and morphometric parameters, often in combination with disease resistance traits such as yellow rust resistance. Advances in molecular breeding have also enabled the development of functional wheat varieties, including those with low gluten content and improved baking quality, as demonstrated by the identification and integration of specific glutenin subunit genes. These achievements contribute to both food security and the development of wheat products tailored to diverse dietary needs.
5 Challenges and Bottlenecks in Wheat Transformation Systems
5.1 Genotype dependency
Genotype dependency remains a major obstacle in wheat transformation, as transformation efficiency varies significantly among cultivars, limiting the broad application of genetic engineering and genome editing. Traditional methods often favor a few model genotypes, such as ‘Fielder’ or ‘Bobwhite’, which are more amenable to transformation, while elite commercial varieties remain recalcitrant. Recent advances have targeted this bottleneck by overexpressing regeneration-related genes such as TaWOX5 and TaCB1, which have been shown to dramatically increase transformation efficiency and reduce genotype dependency, enabling successful transformation across a wider range of wheat varieties and even related cereals. Additionally, morphogenic gene-mediated approaches and optimized Agrobacterium-mediated protocols have further alleviated genotype restrictions, making transformation more accessible for diverse germplasm (Kumar et al., 2019; Johnson et al., 2023).
5.2 Efficiency and stability issues
Despite progress, transformation efficiency and the genetic stability of transgenic progeny remain critical challenges. Factors such as donor plant health, developmental stage, and transformation protocol parameters (e.g., centrifugation, vector design) significantly influence efficiency (Kumar et al., 2019). The use of morphogenic regulators (e.g., GRF4-GIF1, ZmBbm, ZmWus2) and improved tissue culture techniques have led to transformation efficiencies exceeding 50% in some cases and reduced the time required for plant regeneration. However, issues such as transgene silencing, insertion site effects, and variable expression levels persist. Studies have shown that transgene integration can occur at diverse chromosomal locations, with stability and expression sometimes affected by DNA methylation and epigenetic factors. Marker-free transformation strategies and careful selection of promoters and selectable markers are being developed to enhance stability and facilitate regulatory approval (Wang et al., 2016; Shrawat and Armstrong, 2018; Liu et al., 2020).
5.3 Biosafety and public acceptance of transgenic wheat
Biosafety and public acceptance are significant hurdles for the commercial deployment of transgenic wheat. Advances in marker-free transformation and co-transformation strategies have improved the prospects for regulatory approval by reducing the presence of foreign selectable markers. Ecological risk and food safety assessments are ongoing, with studies demonstrating stable inheritance and minimal unintended effects in marker-free lines (Liu et al., 2020). Nonetheless, ethical debates and public skepticism persist, particularly regarding the consumption and environmental impact of genetically modified wheat. Transparent risk assessments, effective communication, and the development of cisgenic or genome-edited lines (which may be perceived as less controversial) are crucial for improving public acceptance and facilitating the adoption of biofortified wheat varieties (Wang et al., 2016; Shrawat and Armstrong, 2018).
6 Future Trends and Challenges
6.1 Key strategies to enhance efficiency and broaden genotype applicability
Screening for recipient materials with broad compatibility is essential to overcome genotype dependency in maize transformation. Recent breakthroughs involve the use of regeneration-promoting genes such as Baby boom (Bbm), Wuschel2 (Wus2), and GRF-GIF fusion proteins, which have significantly increased transformation efficiency and enabled successful transformation across a wider range of maize genotypes, including previously recalcitrant elite lines. For example, the overexpression of Bbm and Wus2, combined with spatial regulation and excision systems, has led to genotype-independent transformation with high-quality, fertile plants and improved transformation frequencies (Lowe et al., 2016). Similarly, GRF-GIF chimeras have been shown to boost regeneration and transformation rates without negative effects on plant growth, further broadening the applicability of transformation protocols. The identification and application of additional developmental regulators, such as ZmWIND1 and Wox2a, continue to expand the range of transformable maize varieties.
6.2 Development of high-throughput and precision transformation systems
The development of high-throughput transformation systems is accelerating maize improvement. Optimized Agrobacterium-mediated protocols, the use of growth regulators, and chemical enhancers like uniconazole have increased transformation frequencies and reduced tissue culture cycles, making year-round, large-scale transformation feasible (Zhao et al., 2002). Innovative delivery methods, such as DNA-coated magnetic nanoparticles for pollen transfection, offer culture-free, genotype-independent transformation, bypassing traditional tissue culture bottlenecks.
Precision genome editing technologies, including CRISPR/Cas9, base editing (ABE/CBE), and prime editing, are being rapidly integrated into maize transformation pipelines, enabling targeted modifications for biofortification traits. High-throughput CRISPR/Cas9 mutagenesis and advanced vector systems have streamlined trait gene identification and editing, while DNA-free editing approaches and protoplast-based systems are being developed to further enhance precision and regulatory acceptance (Liu et al., 2020). The integration of microfluidics and automated inoculation devices is anticipated to further increase throughput and reproducibility in transformation workflows (Kausch et al., 2021).
6.3 Integrating ‘omics + breeding’ to accelerate molecular design breeding
The integration of genomics, transcriptomics, proteomics, and metabolomics with traditional and molecular breeding has revolutionized maize improvement. Systematic gene function analysis through multi-omics approaches enables the identification of key genes, quantitative trait loci (QTLs), and alleles associated with nutritional traits, stress tolerance, and yield (Kaur et al., 2021; Farooqi et al., 2022; Sahito et al., 2024). These insights inform the design of precise gene editing targets and facilitate the development of biofortified maize with enhanced micronutrient content, such as provitamin A, iron, and zinc (Sethi et al., 2023).
High-throughput sequencing and phenotyping, combined with marker-assisted and genomic selection, have established rapid breeding pipelines that accelerate the development and deployment of nutritionally enriched maize varieties (Virk et al., 2021). The use of diverse germplasm and the mining of genetic diversity through omics-driven approaches further expand the pool of beneficial alleles, supporting the creation of resilient and nutrient-rich cultivars (Sethi et al., 2023). Computational biology and network analysis of multi-omics data also provide new perspectives for selecting biomolecules of interest and modeling plant architecture for targeted trait improvement (Jiang et al., 2019).
6.4 Advancing social acceptance and biosafety evaluation
Despite technological advances, regulatory and market challenges persist, especially for non-transgenic, gene-edited materials. While conventional breeding and marker-assisted selection enjoy higher public acceptance, the adoption of genome editing and transgenic approaches faces scrutiny regarding ecological safety and food security. Establishing standardized ecological safety assessment systems and robust policy mechanisms is essential for evaluating the risks and benefits of biofortified maize, particularly as new breeding technologies blur the lines between traditional and genetically modified crops (Palacios-Rojas et al., 2020).
Effective communication, advocacy, and transparent risk assessments are critical for building public trust and facilitating the acceptance of biofortified maize. Mainstreaming biofortification traits into competitive varieties and ensuring their regulatory compliance will be key to scaling up the impact of these innovations globally (Virk et al., 2021).
7 Concluding Remarks
Wheat transformation technology has progressed from early experimental stages to an era marked by high efficiency and precision, largely due to innovations such as the use of regeneration-promoting genes (e.g., TaWOX5, TaCB1, GRF4-GIF1) and optimized transformation protocols. These advances have significantly improved transformation efficiency and reduced genotype dependency, enabling successful genetic modification across a broader range of wheat varieties, including elite cultivars and related species. High-throughput and robust systems, such as Agrobacterium-mediated and particle bombardment methods, now support efficient gene delivery and genome editing, facilitating both functional gene discovery and targeted trait improvement.
Despite these achievements, challenges remain. Transformation efficiency and genotype dependency still limit the routine application of these technologies, especially in recalcitrant or commercially important varieties. Integration with industrial-scale breeding and the translation of laboratory protocols to field-ready, high-performing lines require further optimization and standardization. Additionally, the complexity of wheat’s polyploid genome and the need for precise, stable gene edits continue to pose technical hurdles.
Future efforts should prioritize the integration of multiple technologies-combining advanced transformation systems, genome editing, and high-throughput phenotyping-with interdisciplinary collaboration among molecular biologists, breeders, and bioinformaticians. Such collaboration will accelerate functional gene discovery and the development of new, improved wheat varieties tailored for global food security and sustainable agriculture.
Acknowledgments
Thank you to the anonymous peer review for reading the manuscript and providing constructive suggestions for revisions.
Conflict of Interest Disclosure
The authors affirm that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Abdallah N., Elsharawy H., Abulela H., Thilmony R., Abdelhadi A., and Elarabi N., 2022, Multiplex CRISPR/Cas9-mediated genome editing to address drought tolerance in wheat, GM Crops & Food, 16: 1-17.
https://doi.org/10.1080/21645698.2022.2120313
Cram D., Kulkarni M., Buchwaldt M., Rajagopalan N., Bhowmik P., Rozwadowski K., Parkin I., Sharpe A., and Kagale S., 2019, WheatCRISPR: a web-based guide RNA design tool for CRISPR/Cas9-mediated genome editing in wheat, BMC Plant Biology, 19(1): 474.
https://doi.org/10.1186/s12870-019-2097-z
Fahmy A., Hassanein R., Hashem H., Ibrahim A., Shihy O., and Qaid E., 2018, Developing of transgenic wheat cultivars for improved disease resistance, Journal of Applied Biology and Biotechnology, 6: 31-40.
https://doi.org/10.7324/jabb.2018.60206
Farooqi M., Nawaz G., Wani S., Choudhary J., Rana M., Sah R., Afzal M., Zahra Z., Ganie S., Razzaq A., Reyes V., Mahmoud E., Elansary H., El-Abedin T., and Siddique K., 2022, Recent developments in multi-omics and breeding strategies for abiotic stress tolerance in maize (Zea mays L.), Frontiers in Plant Science, 13: 965878.
https://doi.org/10.3389/fpls.2022.965878
Hayta S., Smedley M., Clarke M., Forner M., and Harwood W., 2021, An efficient Agrobacterium-mediated transformation protocol for hexaploid and tetraploid wheat, Current Protocols, 1(3): e58.
https://doi.org/10.1002/cpz1.58
Hensel G., Marthe C., and Kumlehn J., 2017, Agrobacterium-mediated transformation of wheat using immature embryos, Methods in Molecular Biology, 1679: 129-139.
https://doi.org/10.1007/978-1-4939-7337-8_8
Hu T., Metz S., Chay C., Zhou H., Biest N., Chen G., Cheng M., Feng X., Radionenko M., Lu F., and Fry J., 2003, Agrobacterium-mediated large-scale transformation of wheat (Triticum aestivum L.) using glyphosate selection, Plant Cell Reports, 21: 1010-1019.
https://doi.org/10.1007/s00299-003-0617-6
Jiang J., Xing F., Wang C., Zeng X., and Zou Q., 2019, Investigation and development of maize fused network analysis with multi-omics, Plant Physiology and Biochemistry, 141: 380-387.
https://doi.org/10.1016/j.plaphy.2019.06.016
Johnson K., Chu U., Anthony G., Wu E., Che P., and Jones T., 2023, Rapid and highly efficient morphogenic gene-mediated hexaploid wheat transformation, Frontiers in Plant Science, 14: 1151762.
https://doi.org/10.3389/fpls.2023.1151762
Kamiya Y., Abe F., Mikami M., Endo M., and Kawaura K., 2020, A rapid method for detection of mutations induced by CRISPR/Cas9-based genome editing in common wheat, Plant Biotechnology, 37(2): 247-251.
https://doi.org/10.5511/plantbiotechnology.20.0404b
Kang M., Lee K., Finley T., Chappell H., Veena V., and Wang K., 2022, An improved Agrobacterium-mediated transformation and genome-editing method for maize inbred B104 using a ternary vector system and immature embryos, Frontiers in Plant Science, 13: 860971.
https://doi.org/10.3389/fpls.2022.860971
Kaur B., Sandhu K., Kamal R., Kaur K., Singh J., Röder M., and Muqaddasi Q., 2021, Omics for the improvement of abiotic, biotic, and agronomic traits in major cereal crops: applications, challenges, and prospects, Plants, 10(10): 1989.
https://doi.org/10.3390/plants10101989
Kausch A., Wang K., Kaeppler H., and Gordon-Kamm W., 2021, Maize transformation: history, progress, and perspectives, Molecular Breeding, 41(6): 38.
https://doi.org/10.1007/s11032-021-01225-0
Kim S., Sim J., Lee S., Kim B., Goo T., Lim S., and Lee J., 2021, Establishment of a stable particle bombardment transformation method in Korean commercial wheat (Triticum aestivum L.) varieties, Plant Biotechnology Reports, 15: 151-161.
https://doi.org/10.1007/s11816-021-00670-9
Kumar R., Mamrutha H., Kaur A., Venkatesh K., Grewal A., Kumar R., and Tiwari V., 2017, Development of an efficient and reproducible regeneration system in wheat (Triticum aestivum L.), Physiology and Molecular Biology of Plants, 23: 945-954
https://doi.org/10.1007/s12298-017-0463-6
Kumar R., Mamrutha H., Kaur A., Venkatesh K., Sharma D., and Singh G., 2019, Optimization of Agrobacterium-mediated transformation in spring bread wheat using mature and immature embryos, Molecular Biology Reports, 46: 1845-1853.
https://doi.org/10.1007/s11033-019-04637-6
Li J., Ye X., An B., Du L., and Xu H., 2012, Genetic transformation of wheat: current status and future prospects, Plant Biotechnology Reports, 6: 183-193.
https://doi.org/10.1007/s11816-011-0213-0
Li J., Zhang S., Zhang R., Gao J., Qi Y., Song G., Li W., Li Y., and Li G., 2020, Efficient multiplex genome editing by CRISPR/Cas9 in common wheat, Plant Biotechnology Journal, 19: 427-429.
https://doi.org/10.1111/pbi.13508
Liang Z., Chen K., Zhang Y., Liu J., Yin K., Qiu J., and Gao C., 2018, Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleoproteins, Nature Protocols, 13: 413-430
https://doi.org/10.1038/nprot.2017.145
Liu H., Zhang W., Wang W., Wu J., Wang K., Lin Z., and Ye X., 2022, Identification of three wheat near-isogenic lines originated from CB037 on tissue culture and transformation capacities, Plant Cell, Tissue and Organ Culture, 152: 67-79.
https://doi.org/10.1007/s11240-022-02389-z
Liu H., Wang K., Wang J., Du L., Pei X., and Ye X., 2020, Genetic and agronomic traits stability of marker-free transgenic wheat plants generated from Agrobacterium-mediated co-transformation in T2 and T3 generations, Journal of Integrative Agriculture, 19: 23-32
https://doi.org/10.1016/S2095-3119(19)62601-8
Liu H., Jian L., Xu J., Zhang Q., Zhang M., Jin M., Peng Y., Yan J., Han B., Liu J., Gao F., Liu X., Huang L., Wei W., Ding Y., Yang X., Li Z., Zhang M., Sun J., Bai M., Song W., Chen H., Sun X., Li W., Lu Y., Liu Y., Zhao J., Qian Y., Jackson D., Fernie A., and Yan J., 2020, High-throughput CRISPR/Cas9 mutagenesis streamlines trait gene identification in maize, Plant Cell, 32: 1397-1413.
https://doi.org/10.1105/tpc.19.00934
Lowe K., Wu E., Wang N., Hoerster G., Hastings C., Cho M., Scelonge C., Lenderts B., Chamberlin M., Cushatt J., Wang L., Ryan L., Khan T., Chow-Yiu J., Hua W., Yu M., Banh J., Bao Z., Brink K., Igo E., Rudrappa B., Shamseer P., Bruce W., Newman L., Shen B., Zheng P., Bidney D., Falco C., Register J., Zhao Z., Xu D., Jones T., and Gordon-Kamm W., 2016, Morphogenic regulators Baby boom and Wuschel improve monocot transformation, Plant Cell, 28: 1998-2015.
https://doi.org/10.1105/tpc.16.00124
Palacios-Rojas N., McCulley L., Kaeppler M., Titcomb T., Gunaratna N., Lopez-Ridaura S., and Tanumihardjo S., 2020, Mining maize diversity and improving its nutritional aspects within agro-food systems, Comprehensive Reviews in Food Science and Food Safety, 19(4): 1809-1834.
https://doi.org/10.1111/1541-4337.12552
Qiu D., Hu W., Zhou Y., Xiao J., Hu R., Wei Q., Zhang Y., Feng J., Sun F., Sun J., Yang G., and He G., 2021, TaASR1-D confers abiotic stress resistance by affecting ROS accumulation and ABA signalling in transgenic wheat, Plant Biotechnology Journal, 19: 1588-1601.
https://doi.org/10.1111/pbi.13572
Richardson T., Thistleton J., Higgins T., Howitt C., and Ayliffe M., 2014, Efficient Agrobacterium transformation of elite wheat germplasm without selection, Plant Cell, Tissue and Organ Culture, 119: 647-659.
https://doi.org/10.1007/s11240-014-0564-7
Sahito J., Zhang H., Gishkori Z., Wang Z., Ding D., Zhang X., and Tang J., 2024, Advancements and prospects of genome-wide association studies (GWAS) in maize, International Journal of Molecular Sciences, 25: 1918.
https://doi.org/10.3390/ijms25031918
Sethi M., Saini D., Devi V., Kaur C., Singh M., Singh J., Pruthi G., Kaur A., Singh A., and Chaudhary D., 2023, Unravelling the genetic framework associated with grain quality and yield-related traits in maize (Zea mays L.), Frontiers in Genetics, 14: 1248697.
https://doi.org/10.3389/fgene.2023.1248697
Shrawat A., and Armstrong C., 2018, Development and application of genetic engineering for wheat improvement, Critical Reviews in Plant Sciences, 37: 335-421.
https://doi.org/10.1080/07352689.2018.1514718
Smedley M., Hayta S., Clarke M., and Harwood W., 2021, CRISPR-Cas9-based genome editing in wheat, Current Protocols, 1(3): e65.
https://doi.org/10.1002/cpz1.65
Sparks C., and Doherty A., 2020, Genetic transformation of common wheat (Triticum aestivum L.) using biolistics, Methods in Molecular Biology, 2124: 229-250.
https://doi.org/10.1007/978-1-0716-0356-7_12
Virk P., Andersson M., Arcos J., Govindaraj M., and Pfeiffer W., 2021, Transition from targeted breeding to mainstreaming of biofortification traits in crop improvement programs, Frontiers in Plant Science, 12: 703990.
https://doi.org/10.3389/fpls.2021.703990
Waites J., Mohan V., Achary M., Syombua E., Hearne S., Bandyopadhyay A., Wang D., and Barone P., 2025, CRISPR-mediated genome editing of wheat for enhancing disease resistance, Frontiers in Genome Editing, 7: 1542487.
https://doi.org/10.3389/fgeed.2025.1542487
Wang K., Liu H., Du L., and Ye X., 2016, Generation of marker-free transgenic hexaploid wheat via an Agrobacterium-mediated co-transformation strategy in commercial Chinese wheat varieties, Plant Biotechnology Journal, 15: 614-623.
https://doi.org/10.1111/pbi.12660
Wang W., Pan Q., He F., Akhunova A., Chao S., Trick H., and Akhunov E., 2018, Transgenerational CRISPR-Cas9 activity facilitates multiplex gene editing in allopolyploid wheat, The CRISPR Journal, 1: 65-74.
https://doi.org/10.1089/crispr.2017.0010
Yin G., Wang Y., She M., Du L., Xu H., and Ye X., 2011, Establishment of a highly efficient regeneration system for the mature embryo culture of wheat, Agricultural Sciences in China, 10: 9-17.
https://doi.org/10.1016/S1671-2927(11)60302-7
Zhang Z., Hua L., Gupta A., Tricoli D., Edwards K., Yang B., and Li W., 2019, Development of an Agrobacterium-delivered CRISPR/Cas9 system for wheat genome editing, Plant Biotechnology Journal, 17: 1623-1635.
https://doi.org/10.1111/pbi.13088

. HTML
Associated material
. Readers' comments
Other articles by authors
. Chunxiang Ma
Related articles
. Wheat
. Genetic transformation
. Agrobacterium-mediated transformation
. Gene editing
. Genotype-independent transformation system
Tools
. Post a comment


