Research Insight
Advances in Scomberomorus Genome Evolution-Drivers of Adaptive Evolution and Environmental Selection 
2 Tropical Specialty Crops Research Center, Hainan Institute of Tropical Agricultural Resources, Sanya, 572026, Hainan, China
 Author
Author  Correspondence author
Correspondence author
International Journal of Molecular Ecology and Conservation, 2025, Vol. 15, No. 3
Received: 22 Mar., 2025 Accepted: 29 Apr., 2025 Published: 15 May, 2025
This study systematically analyzes the research progress of Spanish mackerel genome evolution, focusing on its adaptive evolution mechanism and the driving role of environmental selection. Starting from the biologi-cal characteristics and ecological adaptation of the species, the current status of Spanish mackerel genome research is explained, and the molecular mechanism of Spanish mackerel adaptive evolution is explained, such as the molecular adaptation mechanism of Spanish mackerel to environmental factors such as tempera-ture and salinity. The research progress of Spanish mackerel genome evolution driven by environmental se-lection is introduced, and the adaptive evolution examples at the genome level were analyzed with specific cases. Comparative genomics studies have shown that Spanish mackerel is unique in genome structure and function compared with other scombroid fish, which can help identify key genes and signaling pathways re-lated to environmental adaptation. The progress of functional genomics research shows that stress resistance genes such as heat shock proteins are selected and expanded under environmental stresses such as salinity and temperature, while specific metabolic genes (such as FADS2 related to Omega-3 fatty acid synthesis) may participate in the adaptation process through epigenetic regulation. At the same time, this study also summa-rizes the challenges faced by current research and looked forward to future research directions and application prospects, in order to provide a reference for the sustainable utilization and ecological protection of Spanish mackerel resources.
1 Introduction
The Spanish mackerel (commonly known as bā yú) belongs to the family Scombridae and holds significant importance in marine ecosystems and fisheries. The Scombridae family includes about 51 species of pelagic fish, which are widely distributed in coastal and oceanic waters in tropical, subtropical and temperate zones. As a branch of the Scombridae family, Spanish mackerel (genus Scomberomorus) has nearly 20 species, which are mainly distributed along the coasts of tropical to temperate waters around the world, such as the northwest Pacific Ocean, the Indian Ocean and parts of the Atlantic Ocean, and have long-distance migratory habits (Nesnas et al., 2022; Zeng et al., 2022). Spanish mackerel is of moderate size (generally 50 to 100 cm long for adult fish), has delicious meat, is rich in high-quality protein and omega-3 polyunsaturated fatty acids, and is an important source of nutrition in the human diet. Therefore, Spanish mackerel has always been an important economic fish species in coastal fisheries, and high-yield fisheries have been formed in East Asia, Southeast Asia, the Indian Ocean coast and other regions (Shang et al., 2022; Santana et al., 2024). Spanish mackerel is at a high trophic level in the food web and is a typical top predator, mainly preying on small and medium-sized fish and cephalopods. The rise and fall of its population is of great significance to maintaining the stability of the marine ecosystem and the sustainable use of fishery resources.
However, Spanish mackerel resources have faced severe challenges in recent years. On the one hand, overfishing has caused a decline in the number of some populations and changes in life history parameters such as early maturity and miniaturization of individuals; on the other hand, climate warming and changes in the marine environment are affecting its spawning and habitat. For example, rising water temperatures in the northwest Pacific Ocean are believed to have reduced the hatching rate of Japanese Spanish mackerel eggs and the survival rate of juveniles, resulting in limited population replenishment (Yang et al., 2022). Some species of Spanish mackerel have been listed as near-threatened or vulnerable species (such as Australian spotted Spanish mackerel S. munroi, narrow-banded Spanish mackerel S. commerson, etc.), and need to be strengthened in protection (Jiang et al., 2020). Therefore, from the perspective of conservation and fishery management, it is urgent to study the evolutionary potential and mechanism of Spanish mackerel in adapting to environmental changes.
Genome is the basis for understanding the adaptive evolution of species. At present, the research on the biology and population dynamics of Spanish mackerel is mostly focused on fishery assessment, nutritional quality and environmental pollution impact (Gao et al., 2024). Relatively speaking, the research on its phylogenetic relationship, genetic diversity and adaptive evolution mechanism is relatively weak. Genomic research can reveal the evolutionary history and adaptation mechanism of species from the whole genome, which is the key to analyzing how Spanish mackerel achieves local adaptation in a high gene flow environment. High-quality genome sequence and annotation can help determine the functional gene set and provide a platform for comparative genomics and population genomics analysis. At the same time, the acquisition of reference genomes is also crucial for the development of high-throughput molecular markers (such as SNP chips) for population genetic structure analysis and resource population division.
This study will analyze the latest progress in the study of Spanish mackerel genome evolution, focusing on its adaptive evolution mechanism and the role of environmental selection in it. It will introduce the taxonomic status, biological characteristics and ecological adaptability of Spanish mackerel, provide a biological background for genome research, and outline the current research progress in Spanish mackerel genome sequencing and assembly, comparative genomics and functional genomics, and summarize the genome data and findings that have been obtained. Subsequently, the molecular mechanism of Spanish mackerel adaptive evolution is discussed in detail, focusing on how environmental selection drives Spanish mackerel genome evolution, and explaining the impact of environmental factors, the identification method of adaptive gene loci and the selection signals that have been discovered. Through specific cases, the adaptive evolution pattern of Spanish mackerel genome under the scenarios of temperature, salinity changes and migratory behavior is analyzed. This study not only provides a reference for the survival of the species itself, but also is a major help in maintaining the function of marine ecosystems.
2 Analysis of Biological Characteristics and Ccological Adaptability of Spanish mackerel
2.1 Taxonomic status and species distribution of Spanish mackerel
The genus Scomberomorus belongs to the family Scomberidae of the order Scombriformes and is a medium-to-large ocean-dwelling migratory fish. The family Scomberomorus includes more than 50 species, including mackerel, tuna and Spanish mackerel, which are widely distributed in tropical, subtropical and temperate waters around the world. Among them, the genus Scomberomorus is an important branch of the subfamily Scomberinae (or Spanish mackerel) of the family Scomberidae (Alrashada, 2022; Widiasih et al., 2022). According to fish classification and molecular phylogenetic studies, Pacific Spanish mackerel (such as Japanese Spanish mackerel) and Atlantic Spanish mackerel (such as Spanish Spanish mackerel Scomberomorus maculatus) differ in morphology and genetics and can be regarded as different evolutionary lineages. The body of Spanish mackerel is streamlined, with two dorsal fins and a deeply forked caudal fin, which is suitable for fast swimming and chasing prey (Figure 1) (Jeena et al., 2022). Most Spanish mackerels are swimming animals in coastal waters, living in waters with higher water temperatures and abundant bait, and forming migratory paths along the coast and continental shelf waters.
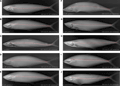 Figure 1 Digital X-ray images of Scomberomorus species (A-J) from the Northern Indian Ocean revealing the diversity, fork length (FL), and number of vertebrae (Adopted from Jeena et al., 2022) |
In terms of geographical distribution, Spanish mackerels are mainly distributed in the coastal waters of the Atlantic, Indian and Pacific Oceans. Among them, the narrow-banded Spanish mackerel (S. commerson) is widely found in the tropical waters of the Indian Ocean and the western Pacific Ocean, from the east coast of Africa through the Red Sea, the Indian subcontinent to the South China Sea and the southern coast of Japan; the Japanese Spanish mackerel (S. niphonius) mainly lives in the temperate waters of the northwest Pacific Ocean, including the Yellow Sea, Bohai Sea and East China Sea of China, and the coast of Japan; while the Atlantic Spanish Spanish mackerel (S. maculatus) is distributed in the warm waters of the western Atlantic Ocean, such as the southeast coast of the United States, the Gulf of Mexico and the Caribbean Sea (Alrashada, 2022; Rothman et al., 2022). As migratory fish, different species of Spanish mackerel often have clear seasonal migration routes and spawning grounds.
2.2 Ecological habits and environmental adaptation characteristics
As a high-speed swimming predatory fish, Spanish mackerel exhibits a series of unique ecological habits and environmental adaptation characteristics. They are usually pelagic or mid-pelagic fish, and are neritic-pelagic species that can move both in the nearshore continental shelf and in the open sea. Take the Japanese mackerel as an example. It is a typical surface-middle layer fish that migrates in the coastal waters of China and surrounding waters throughout the year and has a wide range of tolerance to water temperature and salinity (Muko et al., 2021; Ito et al., 2022). The Indo-Pacific mackerel often appears in coastal waters at a depth of 15 to 200 meters, and prefers turbid, low-salinity nearshore environments such as estuary salt-fresh water mixing areas (Joy et al., 2020; Widiasih et al., 2022). This ability to adapt to lower salinity environments enables it to enter estuaries and bays to feed on prey such as juvenile fish, reflecting a certain degree of euryhaline ecological adaptation.
As a carnivorous fish, Spanish mackerel has swift swimming ability and predation skills. Its streamlined body, well-developed tail fin and muscles give it outstanding explosive power and cruising speed, enabling it to chase fast-moving bait fish. Spanish mackerel feed on small fish such as sardines and anchovies, and also prey on crustaceans and cephalopods such as squid, shrimps and crabs (Fatemi et al., 2017; Sui et al., 2021). The digestive and metabolic systems of Spanish mackerel are also adapted to its predatory and high-speed swimming lifestyle: for example, its muscles are rich in myoglobin and mitochondria to meet the oxygen and energy required for long-term cruising and high-speed sprinting; its tolerance and clearance of lactic acid are also strong, which can quickly restore physical strength after high-intensity exercise.
2.3 Ecological significance of Spanish mackerel adaptive evolution
Understanding the adaptive evolution of Spanish mackerel is of great significance for assessing its ecological status and ability to cope with environmental changes. On the one hand, the evolutionary ability of Spanish mackerel to adapt to a changing environment (such as seasonal changes in water temperature and salinity) determines whether its population can maintain stability in the context of climate change and habitat changes. For example, the rise in water temperature and changes in ocean current patterns in the western Pacific in recent years have had an adverse impact on the reproduction and survival of Japanese Spanish mackerel (Yang et al., 2022). On the other hand, as a high-trophic-level predator, the evolutionary adaptation of Spanish mackerel will also have a chain effect on the downstream ecology-for example, changes in Spanish mackerel predation pressure may affect the succession of pelagic fish populations.
In the context of current global change, research on Spanish mackerel adaptive evolution also has important resource management significance. If genetic variations related to key traits such as stress resistance and fecundity can be identified in the Spanish mackerel genome, it will help predict the potential response of its population to environmental changes (such as rising sea temperatures and changes in salinity). In addition, there may be genetic differentiation and local adaptation between Spanish mackerel populations in different sea areas. If these differences are not understood and unified management is carried out, some hidden populations may be overexploited or neglected. Population genomics studies have shown that even if the overall gene exchange of marine fish is frequent and high gene flow may mask local adaptation signals, some fine genetic structure and adaptive differences may still exist.
3 Research Progress on the Spanish mackerel Genome
3.1 Genome sequencing and assembly research
Thanks to the development of high-throughput sequencing technology, Spanish mackerel genome sequencing has made breakthrough progress in recent years. In 2024, researchers constructed the chromosome-level reference genome of the Spanish mackerel genus for the first time, with the Indo-Pacific Spanish mackerel (Scomberomorus guttatus) as the object. The total size of the genome is about 798 Mb, which is assembled into 24 pseudo-chromosomes, with a Scaffold N50 of 32.9 Mb and a Contig N50 of 8.84 Mb. Gene prediction obtained about 25,886 protein-coding genes. This is the first high-quality genome sequence of the Spanish mackerel genus, laying the foundation for in-depth research on the genome evolution of fish in this genus. Liu et al. (2025) constructed a high-quality reference genome of the Japanese Spanish mackerel (S. niphonius) and explored its population structure in combination with population resequencing data. The acquisition of the Japanese Spanish mackerel genome fills the gap in Spanish mackerel genetic resources in East Asia and creates conditions for comparing the similarities and differences of the genomes of different Spanish mackerel species.
In the process of assembling the Spanish mackerel genome, researchers usually use the strategy of third-generation long-read sequencing combined with Hi-C assisted assembly to ensure the continuity and accuracy of the assembly. For example, in the study of the Indian-Pacific Spanish mackerel genome, PacBio HiFi read-length sequencing was used to obtain high-precision long fragment sequences, and the assembled Contig was anchored to the chromosomes in combination with Hi-C spatial conformation capture, achieving the assembly of 24 chromosomes. It is worth mentioning that through the comparison of high-quality genomes, some evolutionary characteristics of the Spanish mackerel genome can be found. For example, about 36% of the Indian-Pacific Spanish mackerel genome is repeated sequences, and the genome is highly continuous (Bayona-Vásquez et al., 2017; Zheng et al., 2017).
3.2 Progress in comparative genomics
Comparative genomics studies of Spanish mackerel mainly focus on comparing genomic characteristics and evolutionary laws within the Scombridae family and with other pelagic fish. On the one hand, within the Scombridae family, by comparing the genomes of Spanish mackerel with those of tuna, mackerel, etc., shared and unique genomic changes can be identified. For example, Machado et al. (2022) constructed the genome of Atlantic mackerel and pointed out that as an important pelagic fish, it is comparable to Spanish mackerel in terms of genome size and number of coding genes, both containing about 3 billion base pairs and about 30 000 genes. This suggests that the overall genome size and gene number of Scombridae fish are relatively close. However, through fine comparison, it can also be found that some genes have copy number variations or sequence differences between different lineages. For example, Havelka et al. (2021) reported a chromosome-level genome of a closely related species in the Scombridae family-tunas of the genus Euthynnus (such as Euthynnus affinis)-and annotated it using transcriptome data.
On the other hand, comparing the Spanish mackerel genome with pelagic fish with very different environmental adaptations can help reveal the genomic basis of adaptive evolution. For example, tuna have a special adaptation of partial endothermic ability (increasing body temperature), and studies have found that some metabolic and muscle-related genes are expanding or evolving rapidly in the tuna genome (Ciezarek et al., 2020). At present, comparative genomics research on Spanish mackerel is still in its infancy, but existing studies have shown its application prospects. For example, by comparing the genomes of the Indo-Pacific Spanish mackerel with other fish, it was found that some unique repetitive sequences in its genome were suppressed inside tandem repeat genes. For another example, a comparison of the Spanish mackerel and mackerel genomes revealed that many chromosome fragments of the two were colinear, but there were also a few chromosome rearrangements and breaks. These variations may be related to adaptation to different ecological niches during their respective evolution (Li et al., 2025).
3.3 Current status of functional genomics research
Functional genomics focuses on the level of gene expression and gene function, exploring how the genome plays a role in phenotypic adaptation. In the study of Spanish mackerel, functional genomics work mainly includes transcriptome analysis, proteome and epigenetic regulation research. At present, since Spanish mackerel is not a model organism, its functional genomics research is relatively limited, but initial progress has been made. In Spanish mackerel, some studies have begun to focus on the expression changes and selection patterns of functional gene families such as heat shock proteins and immune proteins under environmental stress, as well as the role of epigenetics in local adaptation of populations. For example, Lee et al. (2023) used single-molecule sequencing to simultaneously obtain DNA methylation information during the assembly of the Pacific mackerel genome. In the Indo-Pacific Spanish mackerel genome project, RNA from multiple tissues such as muscle, liver, spleen, and kidney was collected to construct transcriptomes to assist gene annotation. These transcription data not only improve the accuracy of gene prediction, but also lay the foundation for analyzing the expression characteristics of important functional genes.
4 Molecular Mechanisms of Adaptive Evolution of Spanish mackerel
4.1 Key genes related to environmental adaptation
The ability of Spanish mackerel to adapt to the changing marine environment is inseparable from the role of a series of functional genes. The molecular mechanism of its environmental adaptation involves a multi-gene network, including cell stress defense (such as the HSP family), osmotic pressure regulation (ion transporters), energy metabolism and movement (mitochondrial function and muscle protein). At the genomic level, researchers are gradually identifying key genes and gene families related to adaptation to environmental factors such as temperature and salinity. For example, heat shock proteins (HSPs) play an important role in cellular stress response as molecular chaperones. Fish usually regulate salt balance in the body through ion transporters (such as Na^+/K^+-ATPase, ion channel proteins, etc.) expressed in gills and kidneys. Therefore, genes involved in ion transport and osmotic pressure regulation are crucial for Spanish mackerel to adapt to different salinity environments. Some studies have shown that in order to adapt to the extremely cold and low-oxygen environment, the hemoglobin genes of Antarctic fish have changed or even disappeared; in contrast, Spanish mackerel are in a warm and oxygen-rich environment, and may have evolved different blood oxygen transport strategies (Wang et al., 2025).
4.2 Evolutionary mechanisms of immunity and disease resistance
There are many types of pathogens in the marine environment, and Spanish mackerel have also developed unique immune defense mechanisms during the long evolution process. A phylogenetic study of the complement system of bony fish revealed that the complement 3 (C3) and factor H (Cfh) genes of bony fish have been significantly amplified in the genome (Chen et al., 2025). Similarly, the Cfh gene (complement regulatory factor) of fish has also undergone species-specific duplication, which may help to finely regulate complement activity and prevent damage to their own tissues. Therefore, in the Spanish mackerel genome, it is expected that there will also be a phenomenon of complement-related gene amplification, enabling it to fight against a variety of microbial threats in the open ocean environment.
In addition to complement, other immune gene families of fish also reflect adaptive evolutionary characteristics. For example, the antiviral interferon system, antibacterial lysozyme and antimicrobial peptides, and MHC genes responsible for specific immunity have been reported to be expanded or positively selected in different species. Although there are few studies on the specific immune gene evolution of Spanish mackerel, it can be speculated that due to the wide distribution of Spanish mackerel and its migration to different sea areas, it is exposed to a variety of pathogens, and its immune genome may have undergone complex selection pressures.
4.3 Molecular basis of metabolic adaptation mechanisms
Spanish mackerel is at the top of the marine food chain and needs a strong metabolic system to support its high-speed swimming and predation activities. The molecular basis of its metabolic adaptation has been a focus of research in recent years. In fish, there are many genes that affect metabolic rate and environmental tolerance, including enzyme encoding genes involved in energy metabolism pathways, hormone genes that regulate growth and development, and genes that maintain osmotic balance. The metabolic adaptive evolution of Spanish mackerel involves multiple levels, including basal metabolic rate, temperature tolerance, and nutrient utilization (Fauvelot and Borsa, 2011). Its molecular basis is the result of multi-gene coordinated evolution, including amino acid substitution and gene duplication of structural genes, as well as variation in regulatory regions and epigenetic regulation. In the future, by comparing the FADS gene family and its regulatory region sequences of Spanish mackerel and other fish, we may be able to discover genomic differences related to fat metabolism adaptation.
5 Research Progress on Environmental Selection Driving Genome Evolution of Spanish mackerel
5.1 Impact of environmental factors on genome evolution
The diversity and dynamic changes of the marine environment play an important driving role in the genome evolution of marine fish. For Spanish mackerel, which is highly migratory and widely distributed, gradients in temperature, salinity, food resources, etc. in different regions may exert selection pressure on the population. Among them, water temperature gradient is often considered to be one of the key factors driving the adaptive differentiation of marine fish. In the western Pacific, due to the difference in latitude, the Yellow Sea and Bohai Sea in the north and the East China Sea and South China Sea in the south form sea areas with large seasonal temperature differences. For example, in terms of salinity, some Spanish mackerel species (such as the Indo-Pacific Spanish mackerel) prefer low-salinity environments in estuaries, which may cause them to differ in osmotic pressure regulation genes from populations that mainly live in high-salinity waters (Ito et al., 2022; Yang et al., 2022). On the issue of hypoxia, although Spanish mackerel mostly live in surface oxygen-rich waters, if a regional mid-layer hypoxic zone appears in the ocean in the future, its long-term impact may also become a new selection factor.
5.2 Methods for identifying adaptive loci in the genome
Identifying adaptive loci in the genome that are affected by environmental selection is the core task of understanding environmentally driven evolution. Commonly used research methods can be summarized into two categories: detection methods based on population differentiation and detection methods based on environmental association. The first category is the population differentiation method, a typical representative of which is F_ST outlier analysis. The basic idea is to compare the degree of differentiation of different populations at each genetic marker. If the differentiation level of some loci is significantly higher than the background level under the neutral hypothesis, they are regarded as candidate loci affected by selection. This type of method includes the classic F_ST distribution method, Bayesian hierarchical model (such as BayeScan) and principal component analysis (Balding and Beaumont, 2004).
The second category is environmental association analysis, also known as landscape genomics method. Its characteristic is to directly associate genetic variation with environmental variables and find loci where allele frequency is significantly correlated with environmental factors (such as temperature, salinity, etc.). Typical methods include genotype-environment association analysis (GEA), such as Bayenv, LFMM and other models, and spatial analysis based on pointwise regression. Compared with pure population comparison, environmental association analysis can locate specific environmental factors that drive selection (Foll and Gaggiotti, 2008).
5.3 Research progress on selection signals driven by the environment
In species with highly continuous distribution, examples of weak selection signals are often observed. Segovia et al. (2024) showed that in different environments, although the overall genetic homogeneity was high, a very small number of genomic loci were detected to show signs of adaptive differentiation. By analogy to the widespread Spanish mackerel, its adaptive evolutionary signal may also be overwhelmed by a large number of neutral variations. Currently, the public genetic studies of Spanish mackerel populations rarely report significant local adaptation signals, and most believe that its population genetic structure is shallow. For example, Magallon-Gayon et al. (2016) studied the Monterey Spanish mackerel (S. concolor) from the Gulf of California and found that the genetic variation of this species in different regions of the Gulf was highly homogeneous, presenting a global population with random mating.
In recent years, some high-throughput studies have begun to report genomic selection signals in Spanish mackerel and tuna. Feutry et al. (2025) used whole genome scanning to compare different geographical populations of three closely related migratory fishes in the Indian Ocean (including narrow-banded Spanish mackerel), and the results revealed a wide range of genetic differentiation structures. It is worth noting that the anthropogenic environment (such as fishing) also exerts selection on the evolution of Spanish mackerel. Long-term overfishing is believed to have led to the "artificial selection" effect (FIE) of fish, such as overfishing tending to large individuals will select small precocious individuals for survival. For Japanese Spanish mackerel, phenotypic changes such as earlier maturity age and smaller body size have been observed after decades of high-intensity trawling. In the study of adaptive evolution of swordfish, a breakthrough discovery is the role of epigenetics such as DNA methylation in local adaptation. Japanese swordfish research shows that despite the lack of genetic differentiation between populations in different waters along the Chinese coast, their epigenomes differ systematically between the north and the south.
6 Specific Case Studies of Adaptive Evolution at the Genome Level
6.1 Case analysis of temperature adaptation
Temperature is a key environmental factor affecting the geographical distribution and reproductive success of marine organisms. For migratory fish such as Spanish mackerel, temperature adaptability is directly related to their annual migration range and the success or failure of spawning and hatching. The population of Pacific mackerel (similar to Spanish mackerel in ecological niche) showed a population decline under the background of climate warming, which was attributed to the reduction of habitat suitable for spawning and hatching due to rising sea temperatures (Lee et al., 2022; Ray et al., 2022). This shows that the species has not yet accumulated enough high-temperature resistant mutations through evolution to cope with the rapid rise in temperature. Atlantic silverside (Menidia menidia) is a nearshore fish with a wide distribution across latitudes. Studies have found that there are obvious differences in growth and reproduction between northern and southern populations, which are considered to be the result of local adaptation to different temperature environments. Whole-genome analysis further pointed out that there are a large number of linked genomic regions that have undergone differential selection behind these phenotypic differences. Although silverside is not a Spanish mackerel, the "genomic adaptation along the latitudinal gradient" it embodies may also exist in Spanish mackerel.
6.2 Analysis of genomic adaptation under salinity changes
Spanish mackerels usually live in high-salinity marine environments, but some species also regularly enter coastal waters with low salinity, such as estuaries. Salinity fluctuations challenge the osmotic pressure regulation mechanism of fish, and therefore provide an opportunity to study the evolution of salinity adaptation. A typical case comes from the Indo-Pacific Spanish mackerel (S. guttatus). This species often appears in mixed salt waters in estuaries, and the salinity of its environment is lower and fluctuates more than that of a completely marine environment. This suggests that the Indo-Pacific Spanish mackerel has a certain ability to tolerate low salt (Noegroho, 2020).
Although there is no direct report on the salinity adaptation genomics of Spanish mackerel, data from other fish can be used as a reference. For example, in the study of spotted sea bass, it was found that its HSP gene evolved rapidly and was co-expressed under high alkalinity and high salt stress. HSP genes are not only sensitive to temperature stress, but also to salinity stress, and participate in protecting the normal folding function of cell proteins. This shows that fish may partially adapt to salinity changes through the evolution of molecular chaperone systems. Spanish mackerel may also have a similar mechanism: when salinity drops sharply, stress proteins such as HSP are rapidly expressed to protect cell functions; in the long run, the number of genotypes that are more tolerant to salinity fluctuations in Spanish mackerel populations is relatively increasing.
6.3 Cases of genomic adaptation related to migratory behavior
Spanish mackerel are famous for their long-distance migration and seasonal migration. Migratory behavior involves complex physiological and behavioral adaptations, including navigation orientation, long-distance swimming endurance, and gonad development timing. At the genomic level, migration-related traits may be controlled by multiple genes, and current research is beginning to reveal some clues. The differences between the northern and southern populations of Japanese Spanish mackerel provide an interesting case. As mentioned above, the Japanese mackerel along the coast of China can be divided into two groups, the northern and southern groups, which differ in their migration routes and spawning times: the northern group mainly spawns in the Bohai Sea and the Yellow Sea and migrates south to the East China Sea for wintering, while the southern group may reside in the East China Sea and the South China Sea (Yang et al., 2022) (Figure 2). Whole-genome methylation analysis showed that the two groups had different DNA methylation patterns in several genes related to movement and behavior. These genes include genes involved in skeletal muscle development, limb movement (corresponding to the fin and swimming muscle function of fish), and adult individual movement behavior. Differential methylation usually reflects differences in gene expression regulation. This suggests that the northern and southern groups may have phenotypic differences in muscle development, swimming endurance, etc. to adapt to different migration distances and water temperature conditions.
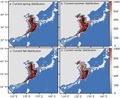 Figure 2 Current simulated habitat and actual survey sites of Scomberomorus niphonius (Adopted from Yang et al., 2022) Image caption: The color gradient indicates variation in habitat suitability (red: highest, blue: lowest). The black dots represent the actual survey sites. (A-D) represent suitable habitats in spring, summer, autumn and winter respectively (Adopted from Yang et al., 2022) |
7 Technical Challenges and Future Directions in the Study of Adaptive Evolution of Spanish mackerel
7.1 Technical challenges in the study
Although the study of Spanish mackerel genome and adaptive evolution has made great progress, it still faces many challenges. The first is the acquisition of high-quality samples and data. Spanish mackerels are mostly oceanic migratory fish, and field sampling and live experiments are difficult, resulting in limited sample size and representativeness of population genomics research. The problem of signal weakening caused by high gene mobility is still prominent. Local adaptation signals of marine fish are often diluted by large-scale gene exchanges. Weak functional verification is also one of the challenges. Although the population genome can indicate which sites are selected, it is difficult to confirm the specific effects of these variations on the phenotype without in-depth functional research. In the context of Spanish mackerel as a non-model species, it is particularly difficult to conduct gene function experiments. Environmental and ecological data acquisition is also a bottleneck. Adaptive evolution research needs to be combined with detailed environmental factor data. However, for highly mobile species such as Spanish mackerel, it is very difficult to track the environmental combinations experienced by individuals. Solving these problems requires multidisciplinary collaboration and technological innovation.
7.2 Future research directions and technology development trends
Currently, there are only a few reference genomes and limited population data for Spanish mackerel species. In the future, it is possible to consider conducting population genome sequencing projects for multiple species of Spanish mackerel, covering representative populations from different geographical regions. Epigenomics will play a more important role in the study of adaptive evolution. The study of DNA methylation in Japanese Spanish mackerel is just a beginning, and it can be expanded to histone modification, non-coding RNA regulation and other aspects in the future. In recent years, environmentally induced heritable epigenetic marks have been considered as a response mechanism of organisms to rapid environmental changes (Gawra et al., 2023). Spanish mackerel has a short generation cycle, and perhaps its adaptation to some environmental pressures is more achieved through epigenetics rather than slow DNA mutations. Therefore, combining genetics and epigenetics for analysis can fully reveal the "genetic traces" of selection.
In addition, functional genomics and experimental evolution will become a trend. With the advancement of gene editing tools, it will be possible to simulate the effects of key gene mutations in Spanish mackerel in the laboratory in the future. The introduction of big data and machine learning will improve analysis efficiency. With the explosive growth of omics data, how to integrate multi-source data such as genome, transcriptome, and environment has become a challenge. In fishery management, establishing a genome-based model to predict the response of populations to future environmental scenarios is also an interdisciplinary frontier direction.
7.3 Application prospects of genome evolution research and ecological protection significance
The study of adaptive evolution of Spanish mackerel is not only of academic value, but also can provide guidance for fishery resource management and marine ecological protection. In fishery management, genome-based population structure identification and genetic diversity monitoring will greatly improve management accuracy. Andersson et al. (2024) emphasized that maintaining genetic diversity of highly abundant species is an important challenge facing future fisheries, which needs to be achieved with the help of population genomics methods. In the field of artificial propagation and aquaculture, the results of genomic research also have application potential. Although Spanish mackerel has not yet been farmed on a large scale, attempts have been made to increase and release and cultivate artificial seedlings in some areas. Genomic data can assist in selecting strains with strong adaptability and fast growth for breeding, and monitor the genetic interaction between released seedlings and wild populations.
Species protection under the background of climate change needs to refer to evolutionary adaptation information. Genomic research can be used to predict the vulnerability of Spanish mackerel to future environments. Dahlke et al. (2020) proposed the concept of "thermal adaptation gap" to assess the risk of fish to climate warming, that is, the bottleneck of sensitivity to temperature at a certain stage of life history. If genomic data can reveal that Spanish mackerel lacks sufficient genetic coping ability in this link, conservation actions should be taken as soon as possible. From the perspective of ecosystem management, maintaining the adaptive potential of top predators such as Spanish mackerel is positive for the entire marine ecology. Whether Spanish mackerel can continue to adapt to changes will affect its predator-prey relationship and competitive relationship with other species, thereby affecting the stability of the ecological network.
Acknowledgments
Thanks to the reviewers for their reading and revision suggestions.
Conflict of Interest Disclosure
The authors affirm that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Alrashada Y., 2022, Growth, mortality and exploitation status of the narrow-barred Spanish mackerel Scomberomorus commerson in Saudi Arabian Gulf waters, Egyptian Journal of Aquatic Biology and Fisheries, 2022: 235121.
https://doi.org/10.21608/ejabf.2022.235121
Andersson L., Bekkevold D., Berg F., Farrell E., Felkel S., Ferreira M., Fuentes-Pardo A., Goodall J., and Pettersson M., 2024, How fish population genomics can promote sustainable fisheries: a road map, Annual Review of Animal Biosciences, 12(1): 1-20.
https://doi.org/10.1146/annurev-animal-021122-102933
Balding D., and Beaumont M., 2004, Identifying adaptive genetic divergence among populations from genome scans, Molecular Ecology, 13(4): 969-980.
https://doi.org/10.1111/j.1365-294X.2004.02125.x
Bayona-Vásquez N., Domínguez-Domínguez O., Díaz-Jaimes P., Uribe-Alcocer M., and Glenn T., 2017, Mitochondrial genomes of the Pacific sierra mackerel Scomberomorus sierra and the Monterey Spanish mackerel Scomberomorus concolor (Perciformes: Scombridae), Conservation Genetics Resources, 10: 471-474.
https://doi.org/10.1007/s12686-017-0851-9
Ciezarek A., Savolainen V., Gardner L., and Block B., 2020, Skeletal muscle and cardiac transcriptomics of a regionally endothermic fish, the Pacific bluefin tuna Thunnus orientalis, BMC Genomics, 21: 642.
https://doi.org/10.1186/s12864-020-07058-z
Chen J., Shao J., Nie L., Fei C., Zhao Z., Wu X., and Zhu T., 2025, Palmitoylation-mediated NLRP3 inflammasome activation in teleosts highlights evolutionary divergence in immune regulation, Zoological Research, 46: 3-14.
https://doi.org/10.24272/j.issn.2095-8137.2024.409
Dahlke F.T., Wohlrab S., Butzin M., and Pörtner H.O., 2020, Thermal bottlenecks in the life cycle define climate vulnerability of fish, Science, 369(6499): 65-70.
Fatemi S., Kaymaram F., Mohammadi G., Hoolihan J., and Niamaimandi N., 2017, Contribution to the feeding habits and reproductive biology of narrow-barred Spanish mackerel Scomberomorus commerson (Lacépède, 1801) (Teleostei: Scombridae) in the northern Persian Gulf, Iranian Journal of Ichthyology, 4: 162-170.
https://doi.org/10.22034/IJI.V4I2.215
Fauvelot C., and Borsa P., 2011, Patterns of genetic isolation in a widely distributed pelagic fish, the narrow-barred Spanish mackerel (Scomberomorus commerson), Biological Journal of the Linnean Society, 104: 886-902.
https://doi.org/10.1111/j.1095-8312.2011.01754.x
Feutry P., Foster S., Grewe P.M., Aulich J., Lansdell M., Clear N., Cooper S., Williams A., Johnson G., Dilrukshi T., Shahid U., Ahusan M., Lestari P., Taufik M., Priatna A., Zamroni A., Usmani H., Farley J., Murua H., Marsac F., and Davies C.R., 2025, Genome scans reveal extensive population structure in three neritic tuna and tuna-like species in the Indian Ocean, ICES Journal of Marine Science, 82(2): fsae162.
https://doi.org/10.1093/icesjms/fsae162
Foll M., and Gaggiotti O., 2008, A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective, Genetics, 180: 977-993.
https://doi.org/10.1534/genetics.108.092221
Gao Y., Liu S., Gutang Q., et al., 2024, Chromosome-level genome assembly of Indo-Pacific king mackerel (Scomberomorus guttatus), Scientific Data, 11(1): 1224.
https://doi.org/10.1038/s41597-024-04110-5
Gawra J., Valdivieso A., Roux F., Laporte M., de Lorgeril J., Gueguen Y., Saccas M., Escoubas J.M., Montagnani C., Destoumieux-Garzón D., Lagarde F., Leroy M.A., Haffner P., Petton B., Cosseau C., Morga B., Dégremont L., Mitta G., Grunau C., and Vidal-Dupiol J., 2023, Epigenetic variations are more substantial than genetic variations in rapid adaptation of oyster to Pacific oyster mortality syndrome, Science Advances, 9(36): eadh8990.
Havelka M., Sawayama E., Saito T., Yoshitake K., Saka D., Ineno T., and Matsubara T., 2021, Chromosome-scale genome assembly and transcriptome assembly of Kawakawa Euthynnus affinis: a tuna-like species, Frontiers in Genetics, 12: 739781.
Ito S., Liu S., Alabia I., Tian Y., Cheng J., and Liu Y., 2022, Development of a prey-predator species distribution model for a large piscivorous fish: a case study for Japanese Spanish mackerel Scomberomorus niphonius and Japanese anchovy Engraulis japonicus, Deep Sea Research Part II: Topical Studies in Oceanography, 207: 105227.
https://doi.org/10.1016/j.dsr2.2022.105227
Jeena N.S., Rahuman S., Roul S.K., Azeez P.A., Vinothkumar R., Manas H.M., Nesnas E.A., Rathinam A.M.M., Surya S., Rohit P., Abdussamad E.M., and Gopalakrishnan A., 2022, Resolved and redeemed: a new fleck to the evolutionary divergence in the genus Scomberomorus Lacepède, 1801 (Scombridae) with cryptic speciation, Frontiers in Marine Science, 9: 888463.
https://doi.org/10.3389/fmars.2022.888463
Jiang T., Ye Z., Yang J., Pan X., Xu B., and Tian Y., 2020, Population connectivity in a highly migratory fish, Japanese Spanish mackerel (Scomberomorus niphonius), along the Chinese coast: implications from otolith chemistry, Fisheries Research, 231: 105690.
https://doi.org/10.1016/j.fishres.2020.105690
Joy L., Basheer V., Ravi C., Paulose S., Singh R., Lal K., Divya P., Mohindra V., and Kumar R., 2020, Microsatellite marker development in Spanish mackerel Scomberomorus commerson using third generation sequencing technology, Molecular Biology Reports, 47: 10005-10014.
https://doi.org/10.1007/s11033-020-05975-6
Lee M., Osuka K., Ito S., Lu Q., Mondal S., and Ray A., 2024, Nonunidirectional habitat changes associated with global climate change: the example of the Indo-Pacific king mackerel (Scomberomorus guttatus) in the Taiwan Strait, Fisheries Oceanography, 34(3): e12718.
https://doi.org/10.1111/fog.12718
Lee Y.H., Abueg L., Kim J.K., Kim Y.W., Fedrigo O., Balacco J., and Kim H., 2023, Chromosome-level genome assembly of chub mackerel (Scomber japonicus) from the Indo-Pacific Ocean, Scientific Data, 10(1): 880.
Li A., Liu S., and Yang J., 2025, Structural characteristics of mitochondrial genomes of two species of mackerel and phylogenetic analysis of Scombridae family, Biomolecules, 15(4): 555.
https://doi.org/10.3390/biom15040555
Liu S., Gao Y., Long X., Li K., Gutang Q., Xie H., Wang J., Tian J., Liang B., Lin J., and Liu W., 2025, A possible more precise management unit delineation based on epigenomic differentiation of a long-distance-migratory marine fish Scomberomorus niphonius, Molecular Ecology Resources, 25(6): e14103.
Machado A.M., Gomes-dos-Santos A., Fonseca M.M., da Fonseca R.R., Veríssimo A., Felício M., Capela R., Alves N., Santos M., Salvador-Caramelo F., Domingues M., Ruivo R., Froufe E., and Castro L.F.C., 2022, A genome assembly of the Atlantic chub mackerel (Scomber colias): a valuable teleost fishing resource, GigaByte, 2022: gigabyte40.
https://doi.org/10.1101/2021.11.19.468211
Muko S., Yoda M., Kurota H., and Ohshimo S., 2021, Fluctuations in distribution and relative abundance of Japanese Spanish mackerel Scomberomorus niphonius in the Yellow Sea, East China Sea and Sea of Japan, Regional Studies in Marine Science, 48: 102057.
https://doi.org/10.1016/j.rsma.2021.102057
Nesnas E., Azeez P., Abdussamad E., Jeena N., Rahuman S., Manas H., Gopalakrishnan A., Surya S., Rohit P., Roul S., Rathinam A., and Vinothkumar R., 2022, Resolved and redeemed: a new fleck to the evolutionary divergence in the genus Scomberomorus Lacepède, 1801 (Scombridae) with cryptic speciation, Frontiers in Marine Science, 9: 888463.
https://doi.org/10.3389/fmars.2022.888463
Noegroho T., 2020, Fishery characteristics of Indo-Pacific king mackerel (Scomberomorus guttatus) in Riau Islands waters (IFMA 711), Indonesia, AACL Bioflux, 13(2): 1179-1189.
Rothman S., Goren M., and Diamant A., 2022, Under the radar: co-introduced monogeneans (Polyopisthocotylea: Gastrocotylinea) of the invasive fish Scomberomorus commerson in the Mediterranean Sea, Parasitology Research, 121: 2275-2293.
https://doi.org/10.1007/s00436-022-07560-1
Santana F., Lessa R., Resende A., and Duponchelle F., 2024, Effects of fishing on the Serra Spanish mackerel (Scomberomorus brasiliensis) in Northeast Brazil, Fisheries Management and Ecology, 31(3): e12688.
https://doi.org/10.1111/fme.12688
Segovia N.I., Coral-Santacruz D., and Haye P.A., 2024, Genetic homogeneity and weak signatures of local adaptation in the marine mussel Mytilus chilensis, Scientific Reports, 14(1): 21081.
Shang Y., Heino M., Sun R., Tian Y., and Sun P., 2022, The effects of selective harvest on Japanese Spanish mackerel (Scomberomorus niphonius) phenotypic evolution, Frontiers in Ecology and Evolution, 10: 844693.
https://doi.org/10.3389/fevo.2022.844693
Sui H., Li Y., Xue Y., Zhang C., Xu B., and Ren Y., 2021, Feeding ecology of Japanese Spanish mackerel (Scomberomorus niphonius) along the eastern coastal waters of China, Acta Oceanologica Sinica, 40: 98-107
https://doi.org/10.1007/s13131-021-1796-0
Wang J., Xie H., Lin J., Li K., Gutang Q., Liang B., Long X., Tian J., Liu W., Liu S., and Gao Y., 2025, A possible more precise management unit delineation based on epigenomic differentiation of a long-distance-migratory marine fish Scomberomorus niphonius, Molecular Ecology Resources, 2025: e14103.
https://doi.org/10.1111/1755-0998.14103
Widiasih D., Nugroho H., Pakpahan S., Widayanti R., and Megarani D., 2022, Revealing Spanish mackerel’s diversity in Indonesia through local commodities in the fish market, Biodiversitas Journal of Biological Diversity, 23(2): 2.
https://doi.org/10.13057/biodiv/d23020
Yang T., Han Z., and Liu X., 2022, Predicting the effects of climate change on the suitable habitat of Japanese Spanish mackerel (Scomberomorus niphonius) based on the species distribution model, Frontiers in Marine Science, 9: 927790.
https://doi.org/10.3389/fmars.2022.927790
Zeng X., Sun C., Li S., Zhang Q., Lao Y., Huang X., and Huang J., 2022, DNA barcoding of Scomberomorus (Scombridae, Actinopterygii) reveals cryptic diversity and misidentifications, ZooKeys, 1135: 157-170.
https://doi.org/10.3897/zookeys.1135.93631
Zheng H., Ouyang L., Li S., Lin Y., Zhang M., Jin Y., and Cheng J., 2017, Complete mitochondrial genome analysis of Japanese Spanish mackerel (Scomberomorus niphonius) with phylogenetic consideration, Mitochondrial DNA Part B: Resources, 2: 56-57.
https://doi.org/10.1080/23802359.2017.1280700

. HTML
Associated material
. Readers' comments
Other articles by authors
. Xuelian Jiang
. Rudi Mai
Related articles
. Spanish mackerel ( Scomberomorus )
. Genome
. Adaptive evolution
. Environmental selection
. Comparative geno
Tools
. Post a comment


