 Author
Author  Correspondence author
Correspondence author
International Journal of Molecular Ecology and Conservation, 2024, Vol. 14, No. 6
Received: 01 Oct., 2024 Accepted: 11 Nov., 2024 Published: 05 Dec., 2024
This study summarizes the current status of crustacean biogeography research and explores the integration of big data resources (such as modern and historical distribution records, paleoclimate and paleoecological data, and molecular systematics information) and their application in spatial modeling and evolutionary biogeography research. The study found that historical climate change, sea level change, and continental drift significantly affected the global distribution pattern, community succession, and population dynamics of crustaceans; crustaceans in freshwater, marine, and island environments showed specific response mechanisms. At the same time, this study also pointed out the challenges of data sparsity, time scale asymmetry, data bias, and uncertainty assessment of multi-data fusion in current research. In the future, cross-disciplinary and multi-scale data integration and model optimization should be strengthened to more accurately predict the response of crustaceans to future environmental changes. This study emphasizes that the widespread application of big data has promoted biogeographic research from static to dynamic, from local to global perspectives, and deeply revealed the ecological adaptation and evolution mechanism of crustaceans, providing an important theoretical basis for biodiversity conservation and ecosystem management.
1 Introduction
Crustaceans are a varied group with great importance in nature. For a long time, they have been used as example creatures to learn how environmental changes affect water-based ecosystems. How they change in their body functions, actions, and where they live—both in the past and now—gives key information about how healthy ecosystems are and how well they can bounce back (Pillay, 2019; Marin and Tiunov, 2023; Nozarpour et al., 2023).
Crustaceans are key players in the food chain. They are predators, prey, and decomposers. This multifaceted role drives nutrient flows and sustains ecosystems in the oceans, freshwater, and at the interface of rivers and seas. They are extremely sensitive to environmental stresses—whether it is temperature fluctuations, salinity changes, or pollution—making them natural indicators of ecosystem change and the impacts of human activities and climate (Moullac and Haffner, 2000; Shields, 2019; Lam-Gordillo et al., 2023). Crustaceans are also economically and culturally important, supporting fisheries and aquaculture worldwide (Behringer and Duermit-Moreau, 2020; Azra et al., 2022).
New breakthroughs in habitat research are powered by massive amounts of data. Long-term natural observations, physiological function studies, and molecular assays are being combined to unravel the complex mechanisms by which crustaceans respond to environmental change. More research is beginning to focus on the effects of various stressors—climate change, ocean acidification, habitat alteration, and more—on crustaceans at different scales. These studies are sketching out a detailed picture of adaptation pathways, vulnerable areas, and recovery patterns of crustacean populations (Whiteley, 2011; Shields, 2019; Stein and Harzsch, 2020; Jaffer et al., 2024). Rapid processing of massive amounts of data and cross-disciplinary collaboration are deepening our understanding of how environmental changes (both new and old) shape crustacean habitats and ecosystem operations (Stein and Harzsch, 2020; Azra et al., 2022; Lam-Gordillo et al., 2023).
This study will summarize existing knowledge on the responses of crustaceans to historical environmental change, emphasize the integration of big data in biogeographic analysis, summarize emerging research trends and methodological advances, identify knowledge gaps, and explore future directions for big data to inform the conservation and management of crustacean biodiversity in a changing world. This study provides a reference for exploring the ecological responses of crustaceans to historical climate and environmental changes.
2 Current Status and Challenges in Crustacean Biogeography Research
2.1 Overview of crustacean biogeographic patterns
The biogeographic distribution patterns of crustaceans are rich and diverse, and ecological processes and historical events jointly outline these pictures. In the ocean, planktonic crustaceans are deeply affected by ocean currents, water depth changes and temperature gradients; benthic species are more restricted by continental barriers and seafloor topography. Life history characteristics - such as whether there is a free-swimming larval stage - play a key role in their dispersal ability and local distribution. Decapod species with a swimming larval stage tend to have a wider distribution range; while amphipod species that lack this stage are more likely to be confined to small areas and show significant species replacement between different geographical regions.
The Coral Triangle and the Mediterranean are hotspots of biodiversity with a wide variety of species, including many endemic species. The spatial distribution of benthic crustaceans is not only driven by the environment, but is also clearly constrained by uneven sampling. Taking the northwest Pacific as an example, there are differences in the distribution of biodiversity: benthic species are most densely distributed along the coasts of China and the Philippines, while pelagic species are concentrated in northeastern Japan and the Sea of Okhotsk (Figure 1) (Knauber et al., 2023). In addition, species complexes and cryptic species (species with similar appearance but different genes) make the interpretation of global distribution patterns more complicated. This also highlights the important value of molecular tools in improving classification systems and distribution data (Ghazoul, 2016; Halsband et al., 2020; Arfianti et al., 2020; Riehl and Kühn, 2020; Arfianti and Costello, 2021).
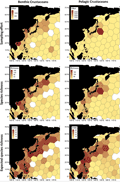 Figure 1 Biodiversity patterns of benthic and pelagic crustaceans in the NWP (Adopted from Knauber et al., 2023) Image caption: Sampling effort (number of distribution records), alpha species richness (number of species per hexagon) and ES50 (expected number of species) of benthic (n = 6939) and pelagic crustaceans (n = 8612) in the NWP is visualized per hexagonal cells (700,000 km2 per cell). Areas without colouring had zero values. Lower counts of coloured grid cells in species richness and expected species richness than in sampling effort are caused by distribution records that lack taxonomic identification to species leve (Adopted from Knauber et al., 2023) |
2.2 Research advances on the impacts of historical environmental changes on crustaceans
In recent years, researchers have used large-scale species distribution records and molecular data to explore how historical environmental changes (such as population isolation events and climate fluctuations) shape crustacean populations. Genetic tests have found that some important fishery species have independent subpopulations, which contrasts with previous understanding of their reproduction and spread methods. Comparative phylogeographic studies across regions have shown that habitat type and life history traits affect the timing and extent of population differentiation: some species show clear genetic differentiation in biogeographic barrier areas, while others do not. These findings not only reveal the complexity of crustacean responses to historical changes, but also emphasize the need to integrate genetic, ecological and distribution data (Arfianti et al., 2020; Ewers-Saucedo and Wares, 2020; Riehl and Kühn, 2020; Peart and Schnabel, 2024).
2.3 Technical challenges and limitations in current studies
Despite progress in this field, crustacean biogeography research still faces several technical challenges. Inaccurate species identification, outdated classification systems, and lack of comprehensive identification guidelines are obstacles to the accurate mapping of distribution maps, especially in the study of important fishery species and cryptic species. There are obvious biases in global sampling, and data from less studied regions are scarce, resulting in incomplete global data sets. Many studies rely on limited samples or lack of repeated verification data, which undermines the reliability of the conclusions. The application of molecular tools is still in its infancy, and there is an urgent need to establish standardized methods and strengthen taxonomic training. Addressing these challenges will be of great significance for improving the resolution of biogeographic patterns and developing more effective conservation and management strategies (Havird and Santos, 2016; Arfianti et al., 2020; Ewers-Saucedo and Wares, 2020; Halsband et al., 2020; Riehl and Kühn, 2020; Yuan et al., 2023).
3 Integration and Application of Big Data Resources in Crustacean Biogeographic Research
3.1 Overview of multisource data types
Today's crustacean biogeography research is increasingly integrating multiple data types in order to fully grasp its current and past distribution patterns. Public databases such as the Global Biodiversity Information Platform (GBIF) and the Marine Biodiversity Information System (OBIS) contain massive standardized species distribution records. These data support researchers to conduct large-scale analyses on a spatiotemporal scale to explore species numbers, distribution patterns, and the environmental drivers behind them - studies on deep-sea crustaceans in the northwest Pacific and Bering Sea are examples (Knauber et al., 2022). Although not directly cited here, paleoclimate data and marine sedimentary records from PaleoClim and PMIP are also indispensable in interpreting historical biogeographic changes.
Fossil records and stable isotope analysis (such as carbon and nitrogen isotopes) are increasingly used in reconstructing the distribution of ancient ecosystems and food web structures. For example, stable isotope data have been used to simulate trophic biogeographic changes in invasive crustaceans, providing support for revealing the succession process of historical ecosystems (Di Muri et al., 2023). Advances in genomics have spawned platforms such as CRUSTADB, which integrates genome assembly, resequencing data, transcriptome and mitochondrial genome information to provide support for phylogenetic comparison, population genomics and evolutionary analysis, making biogeographic research more accurate (Wang et al., 2025).
3.2 Data integration platforms and analytical tools
To make good use of big data in crustacean biogeographic research, powerful analysis platforms and modeling tools are indispensable. Software such as MaxEnt, BioMod, and SDMtoolbox are widely used to construct species distribution models. With environmental variables and species occurrence records, the current and future distribution ranges of species can be predicted (Knauber et al., 2022). Tools such as BEAST and BioGeoBEARS help reconstruct historical biogeographic scenes and evolutionary associations by integrating molecular data and distribution information.
Programming environments such as R and Python, combined with cloud platforms such as Google Earth Engine, have become core tools for processing, analyzing, and visualizing large-scale complex data. Virtual research environments such as the Tesseract platform developed by LifeWatch ERIC provide a user-friendly interface to integrate analysis processes, including stable isotope modeling and spatial analysis (Di Muri et al., 2023; Wang et al., 2025). Together, these tools are driving the widespread application and in-depth development of big data in crustacean biogeography research.
4 Advances in Crustacean Historical Environmental Response Research Driven by Big Data
4.1 Studies on the impact of historical climate change on crustacean distribution patterns
Large-scale physiological and ecological data sets allow researchers to more accurately explain the role of environmental stress (such as temperature fluctuations and changes in oxygen concentration) on the physiological functions, immune levels and distribution patterns of crustaceans through meta-analysis and systematic reviews. For example, meta-analysis of decapod crustaceans has locked in key physiological indicators such as lactate (L-lactate) - these indicators can effectively mark the impact of climate-driven environmental changes on their health and distribution (Conneely and Coates, 2023). In addition, research on the neuro-endocrine-immune regulatory mechanisms of crustaceans has shown that environmental stress (especially stress related to climate change) can directly suppress their immune defenses, thereby threatening individual survival and population continuity (Tong et al., 2021). These advances provide a key basis for understanding how historical and current climate change shapes the distribution of crustaceans.
4.2 Research on sea-level change and crustacean community succession
In the existing literature, there is a lack of research that directly uses big data to explore the relationship between sea level change and crustacean community succession. However, a systematic review of metal migration and toxicity in the marine environment shows that crustaceans are ideal bioindicators for tracking changes in aquatic ecosystems. This type of research uses large-scale data sets to analyze how abiotic factors such as pH, dissolved oxygen, and salinity (whose changes are often related to sea level fluctuations) regulate the bioavailability and toxicity of metals, thereby affecting the health of crustacean populations (De Almeida Rodrigues et al., 2021). This type of research method provides a reference path for exploring the impact of historical sea level fluctuations on the structure and succession dynamics of crustacean communities.
4.3 Continental drift and the evolution of crustacean biogeographic patterns
The papers looked at don't directly talk about continental drift. But progress in molecular systematics and population genomics-made easier by big data platforms-is being used more to rebuild the evolutionary histories and biogeographic patterns of crustaceans. These methods help make clear how old geological events, like continental drift, have helped crustacean groups become more diverse and get to where they are now. This is done by bringing together genetic, ecological, and distribution data across large spaces and long times (Tong et al., 2021).
5 Case Studies on Crustacean Responses to Historical Environmental Changes
5.1 Glacial–interglacial response patterns in freshwater crustaceans
In existing studies, there are not many cases that directly analyze the response of freshwater crustaceans during glacial and interglacial cycles. However, surveys of large benthic crustaceans in estuaries show that climate-related factors - such as sea surface temperature and large climate indices (such as the Southern Oscillation Index) - have a significant impact on the abundance and species diversity of crustaceans in different regions. These findings suggest that historical climate fluctuations similar to glacial-interglacial periods may have largely shaped the distribution range and community structure of freshwater and estuarine crustaceans. This shaping is mainly achieved by changing temperature patterns and water system connectivity (Lam-Gordillo et al., 2023).
5.2 Marine crustacean responses to ancient oceanic climate change
Faced with ancient and contemporary marine climate changes, marine crustaceans have shown obvious responses in both morphology and ecology. For example, studies have found that high sea surface temperatures and ocean acidification—environments similar to ancient marine climate change—can seriously impair the health, immune function, and exoskeleton performance of crustaceans. Meta-analysis shows that high pCO₂ concentrations reduce the calcium and magnesium content of their exoskeletons, causing the shell to become thinner and more brittle, hindering growth and molting, especially for species with weaker regulatory abilities. These changes are directly related to the survival, distribution, and abundance of species, indicating that some marine crustaceans may be in a state of high vulnerability to both ancient and future marine climate changes (Whiteley, 2011; Shields, 2019; Siegel et al., 2022).
5.3 Island crustacean response mechanisms to sea-level fluctuations
There is a lack of detailed case studies on island crustaceans in the current literature, but studies on estuarine and coastal crustaceans provide useful references. Sea level changes—especially climate-related fluctuations—can change sediment composition, salinity levels, and habitat availability, thereby affecting the structure and succession of crustacean communities. For example, increased sedimentation and salinity changes caused by sea level rise have been shown to lead to a decrease in crustacean species and abundance, meaning that island and coastal crustacean communities are quite sensitive to such environmental changes (Lam-Gordillo et al., 2023).
6 Challenges and Frontier Issues
6.1 Data sparsity and temporal scale asymmetry
Crustacean biogeography studies are often plagued by data scarcity, especially in the poorly studied Indian Ocean and some tropical regions. Many taxa - including species that are critical to fisheries and ecology - have not been adequately sampled or scientifically described, resulting in gaps in knowledge about their distribution ranges and historical dynamics. Data imbalances in time scales are also prominent: modern records are often detailed, but historical data are relatively scarce, making it difficult to reconstruct long-term distribution patterns and environmental response processes of crustaceans (Marrone et al., 2017; Arfianti et al., 2020; Arfianti and Costello, 2021; Halsband et al., 2020).
6.2 Effects of modern distribution bias and incomplete specimen records on model accuracy
Sampling isn't done evenly. Most records come from places that have been studied a lot and from middle latitudes, while big areas are left out. This lopsided sampling gives wrong ideas about how many species there are, which ones are found only in certain places, and how species change from one area to another. It can also mess up the results of spatial models. Old or incomplete records of specimens, along with not being able to identify species properly and missing voucher specimens, make biogeographic models and conservation checks less accurate. Not having good, up-to-date guides to identify species and not enough experts in naming species make these problems worse—especially for species that look alike or are hard to tell apart (Arfianti et al., 2020; Arfianti and Costello, 2021; Halsband et al., 2020).
6.3 Uncertainty assessment and methodological improvements in multi-source data integration
When integrating multi-source data such as molecular, habitat and distribution, methodological challenges emerge in an endless stream. Differences in data quality, spatiotemporal accuracy, and inconsistent species naming all introduce uncertainty into the analysis. Although emerging molecular technologies and big data platforms can improve the accuracy of species identification and data precision to a certain extent, standardized integration processes and effective uncertainty assessment methods are still urgently needed. Advanced data analysis methods, such as integration tools in virtual scientific research environments, can alleviate some of these problems, but continuous optimization is still needed in the future to enhance the consistency of data integration and the reproducibility of crustacean biogeography research (Marrone et al., 2017; Halsband et al., 2020; Di Muri et al., 2023).
7 Concluding Remarks
Scientists have been able to look again at and improve what we know about how many crustacean species there are and where they live, across large areas and long times, by using open-access biodiversity databases like OBIS and GBIF. For example, studies of crustaceans in the Northwest Pacific have shown clear differences in the number of species between those that live on the sea floor and those that swim in open water. They have also found key environmental factors, such as how much food is made by plants and salt levels, that affect these patterns. These findings show how useful big data is in finding complex biogeographic patterns and guessing how different crustacean groups might react to climate changes in the future. In the same way, putting deep-sea crustacean data from the Bering Sea into digital form and sharing it is filling important gaps in what we know and helping with global checks on biodiversity.
The way crustaceans have responded to past environmental changes—such as the passing of ice ages, fluctuations in ocean climate, and rising and falling sea levels—provides an excellent example for understanding species evolution and adaptation. Stable isotope detection and DNA barcoding technologies have opened up new perspectives for interpreting food web dynamics, distinguishing species, and discovering hidden differences between species. This confirms the value of cross-disciplinary collaboration in revealing recent and ancient events related to species habitats.
Despite these advances, problems remain: data scarcity, uneven sampling, and the need to improve the integration of molecular, ecological, and distributional data. Building easy-to-use analysis platforms and virtual research spaces can help alleviate these challenges. But greater breakthroughs will rely on deepening model integration and cross-disciplinary collaboration. Only by establishing collaborations between species nomenclature researchers, ecologists, data experts, and policymakers can the field move towards more solid, predictive, and practical biogeographic research.
![]() Acknowledgments
Acknowledgments
I sincerely thank the anonymous peer reviewers for their valuable comments and detailed suggestions..
Conflict of Interest Disclosure
The author affirms that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Arfianti T., Arfianti T., and Costello M., 2020, Global biogeography of marine amphipod crustaceans: latitude, regionalization, and beta diversity, Marine Ecology Progress Series, 638: 83-94.
https://doi.org/10.3354/meps13272
Arfianti T., and Costello M., 2021, The distribution of benthic amphipod crustaceans in Indonesian seas, PeerJ, 9: e12054.
https://doi.org/10.7717/peerj.12054
Azra M., Noor M., Eales J., Sung Y., and Ghaffar M., 2022, What evidence exists for the impact of climate change on the physiology and behaviour of important aquaculture marine crustacean species in Asia? A systematic map protocol, Environmental Evidence, 11(1): 9.
https://doi.org/10.1186/s13750-022-00263-1
Behringer D., and Duermit-Moreau E., 2020, Crustaceans, One Health and the changing ocean, Journal of Invertebrate Pathology, 186: 107500.
https://doi.org/10.1016/j.jip.2020.107500
Conneely E., and Coates C., 2023, Meta-analytic assessment of physiological markers for decapod crustacean welfare, Fish and Fisheries, 25(1): 134-150.
https://doi.org/10.1111/faf.12798
De Almeida Rodrigues P., Ferrari R., Kato L., Hauser-Davis R., and Conte-Junior C., 2021, A systematic review on metal dynamics and marine toxicity risk assessment using crustaceans as bioindicators, Biological Trace Element Research, 200: 881-903.
https://doi.org/10.1007/s12011-021-02685-3
Di Muri C., Arvanitidis C., Basset A., De Giorgi R., Rosati I., Vaira L., and Mancinelli G., 2023, The validation case on invasive crustaceans of the LifeWatch ERIC Internal Joint Initiative: state of the art and next steps forward, Frontiers in Environmental Science, 10: 1038635.
https://doi.org/10.3389/fenvs.2022.1038635
Ewers-Saucedo C., and Wares J., 2020, Population connectivity and phylogeography of crustaceans, Evolution and Biogeography, 8(8): 440-463.
https://doi.org/10.1093/oso/9780190637842.003.0017
Ghazoul J., 2016, Evolution and biogeography, Dipterocarp Biology, Ecology, and Conservation, 2016: 69-88.
https://doi.org/10.1093/acprof:oso/9780199639656.003.0004
Halsband C., Ahyong S., Brandt A., Kosobokova K., Ward P., Goodall-Copestake W., and Macpherson E., 2020, Biogeography of the oceans, The Natural History of the Crustacea, 2020: 121-154.
https://doi.org/10.1093/oso/9780190637842.003.0006
Havird J., and Santos S., 2016, Here we are, but where do we go? A systematic review of crustacean transcriptomic studies from 2014–2015, Integrative and Comparative Biology, 56(6): 1055-1066.
https://doi.org/10.1093/icb/icw061
Jaffer Y., Bhat I., Mir I., Bhat R., Sidiq M., and Jana P., 2024, Adaptation of cultured decapod crustaceans to changing salinities: physiological responses, molecular mechanisms and disease implications, Reviews in Aquaculture, 16(4): 1520-1543.
https://doi.org/10.1111/raq.12909
Knauber H., Kohlenbach K., Böhm P., Lüter C., Ziegler A., Brandt A., and Saeedi H., 2023, Deep-sea benthic crustacean and annelid data from the Bering Sea, Data in Brief, 48: 109186.
https://doi.org/10.1016/j.dib.2023.109186
Knauber H., Kohlenbach K., Brandt A., and Saeedi H., 2022, Crustaceans of the Northwest Pacific Ocean: species richness and distribution patterns, Journal of Sea Research, 191: 102332.
https://doi.org/10.1016/j.seares.2022.102332
Lam-Gordillo O., Lohrer A., Douglas E., Hailes S., Carter K., and Greenfield B., 2023, Scale-dependent influence of multiple environmental drivers on estuarine macrobenthic crustaceans, Frontiers in Marine Science, 10: 1292849.
https://doi.org/10.3389/fmars.2023.1292849
Marin I., and Tiunov A., 2023, Terrestrial crustaceans (Arthropoda, Crustacea): taxonomic diversity, terrestrial adaptations, and ecological functions, ZooKeys, 1169: 95-162.
https://doi.org/10.3897/zookeys.1169.97812
Marrone F., Rogers C., Zarattini P., and Naselli-Flores L., 2017, New challenges in anostracan research: old issues, new perspectives and hot topics, Hydrobiologia, 801: 179-185.
https://doi.org/10.1007/s10750-017-3345-6
Moullac G., and Haffner P., 2000, Environmental factors affecting immune responses in Crustacea, Aquaculture, 191: 121-131.
https://doi.org/10.1016/S0044-8486(00)00422-1
Nozarpour R., Shojaei M., Naderloo R., and Nasi F., 2023, Crustaceans functional diversity in mangroves and adjacent mudflats of the Persian Gulf and Gulf of Oman, Marine Environmental Research, 186: 105919.
https://doi.org/10.1016/j.marenvres.2023.105919
Peart R., and Schnabel K., 2024, Editorial: advances in crustacean research from the 10th International Crustacean Congress, Frontiers in Marine Science, 11: 1529867.
https://doi.org/10.3389/fmars.2024.1529867
Pillay D., 2019, Ecosystem engineering by thalassinidean crustaceans: response variability, contextual dependencies and perspectives on future research, Diversity, 11(4): 64.
https://doi.org/10.3390/d11040064
Riehl T., and Kühn M., 2020, Uniting what belongs together—reevaluation of the isopod species Macrostylis grandis and M. ovata using ontogenetic, morphological and genetic evidence, Progress in Oceanography, 181: 102238.
https://doi.org/10.1016/j.pocean.2019.102238
Siegel K., Kaur M., Grigal A., Metzler R., and Dickinson G., 2022, Meta-analysis suggests negative, but pCO₂-specific, effects of ocean acidification on the structural and functional properties of crustacean biomaterials, Ecology and Evolution, 12(6): e8922.
https://doi.org/10.1002/ece3.8922
Shields J., 2019, Climate change enhances disease processes in crustaceans: case studies in lobsters, crabs, and shrimps, Journal of Crustacean Biology, 39(6): 673-683.
https://doi.org/10.1093/jcbiol/ruz072
Stein W., and Harzsch S., 2020, The neurobiology of ocean change—insights from decapod crustaceans, Zoology, 144: 125887.
https://doi.org/10.1016/j.zool.2020.125887
Tong R., Pan L., Zhang X., and Li Y., 2021, Neuroendocrine-immune regulation mechanism in crustaceans: a review, Reviews in Aquaculture, 14(1): 378-398.
https://doi.org/10.1111/raq.12603
Wang Q., Lv J., Liu P., Ren X., Li J., Li Y., and Li J., 2025, CRUSTADB: an integrated genomics platform for crustaceans, The Innovation Life, 3(1): 100116-1.
https://doi.org/10.59717/j.xinn-life.2024.100116
Whiteley N., 2011, Physiological and ecological responses of crustaceans to ocean acidification, Marine Ecology Progress Series, 430: 257-271.
https://doi.org/10.3354/meps09185
Yuan J., Yu Y., Zhang X., Li S., Xiang J., and Li F., 2023, Recent advances in crustacean genomics and their potential application in aquaculture, Reviews in Aquaculture, 15(4): 1501-1521.
https://doi.org/10.1111/raq.12791

. HTML
Associated material
. Readers' comments
Other articles by authors
. Zhen Li
Related articles
. Crustaceans
. Big data
. Biogeography
. Historical environmental change
. Ecological response
Tools
. Post a comment


