Review Article
Phylogenetic Evolution of the Water Buffalo (Bubalus bubalis): A Comprehensive Review of Molecular Evidence 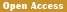
 Author
Author  Correspondence author
Correspondence author
International Journal of Molecular Evolution and Biodiversity, 2024, Vol. 14, No. 6
Received: 26 Sep., 2024 Accepted: 07 Nov., 2024 Published: 23 Nov., 2024
This study presents a comprehensive analysis of the phylogenetic relationships and evolutionary trajectories of river and swamp buffaloes (Bubalus bubalis) through molecular evidence, incorporating mitochondrial DNA (mtDNA), nuclear DNA, and whole-genome sequencing data. The findings suggest that river and swamp buffaloes underwent distinct domestication events in the Indian subcontinent and the China-Southeast Asia region, respectively, with subsequent hybridization events contributing to enhanced genetic diversity among populations. Advances in genomic technologies have provided critical insights, including the identification of a LINE-1 transposon insertion in the ASIP gene, which is responsible for the white coat phenotype in swamp buffaloes. Furthermore, the integration of archaeological and ecological data offers novel perspectives for phylogenetic studies, particularly in addressing challenges such as incomplete lineage sorting (ILS) and sampling bias. By synthesizing molecular, archaeological, and ecological evidence, this study aims to establish a robust phylogenetic framework for water buffaloes, facilitating a deeper understanding of their evolutionary history and genetic diversity.
1 Introduction
The phylogenetic study of the water buffalo (Bubalus bubalis) is fundamental to understanding its evolutionary history and genetic diversity, as it represents an economically and culturally significant livestock species. Water buffaloes are classified into two major types: river buffaloes and swamp buffaloes, each exhibiting distinct genetic and phenotypic characteristics (Kierstein et al., 2004; Curaudeau et al., 2021). The domestication and evolutionary history of these two types have been extensively investigated, yet remain subjects of ongoing discussion and refinement. Integrating molecular, archaeological, and ecological perspectives is essential for resolving uncertainties in their phylogenetic relationships and for constructing a comprehensive evolutionary framework. Molecular evidence, including mitochondrial DNA (mtDNA) and nuclear DNA analyses, has provided insights into the evolutionary pathways and domestication processes of these animals (Mishra et al., 2015; Nagarajan et al., 2015). Understanding the phylogeny of water buffaloes not only sheds light on their evolutionary history but also informs breeding programs and conservation strategies (Mintoo et al., 2019).
Water buffalo play a crucial role in agriculture and ecosystems, particularly across Asia, where they are indispensable for dairy and meat production, directly supporting the livelihoods of millions of people. The river buffalo, predominantly found in the Indian subcontinent and Mediterranean regions, is highly valued for its superior milk yield, whereas the swamp buffalo, common in Southeast Asia and China, is primarily utilized for its draught power and adaptability to wetland environments (Curaudeau et al., 2021). The genetic diversity within and between these two types is critical for their adaptation to diverse environmental conditions and for enhancing economically important traits such as disease resistance and productivity (Vijh et al., 2008). In addition to their agricultural significance, water buffaloes contribute to the maintenance of wetland ecosystems by facilitating nutrient cycling and creating habitats, thereby playing a key role in ecological sustainability (Zhang et al., 2020).
This study seeks to analyze molecular evidence related to the phylogenetic evolution of water buffalo. By synthesizing findings from mitochondrial DNA (mtDNA), nuclear DNA, and microsatellite markers, it aims to elucidate the domestication processes, genetic diversity, and evolutionary relationships between river and swamp buffaloes. This comprehensive examination of the genetic structure and phylogeography of Bubalus bubalis will provide valuable insights for breeding programs, conservation efforts, and the management of both domesticated and wild populations.
2 Taxonomy and Classification of Water Buffalo
2.1 Historical classification and taxonomy
The classification and taxonomy of water buffalo have undergone significant refinements over time, initially based on morphological characteristics and geographical distribution (Sikdar et al., 2021). Traditionally, domestic water buffaloes have been categorized into two major types: the river buffalo and the swamp buffalo, distinguished by their distinct habitats, physical attributes, and utility in agricultural systems. River buffaloes are generally characterized by a black coat and prominently curved horns, whereas swamp buffaloes exhibit a dark grey body, often marked by white chevrons on the throat, white socks, a tail tip, and relatively straight, occasionally elongated horns (Figure 1) (Zhang et al., 2020). These morphological distinctions, alongside genetic analyses, have provided the foundation for a more precise taxonomic framework that continues to evolve with advances in molecular research. River buffaloes are typically found in the Indian subcontinent and Mediterranean countries, while swamp buffaloes are prevalent in China and Southeast Asia (Curaudeau et al., 2021).
.png) Figure 1 Morphological variation in Asian water buffalo breeds (Adopted from Zhang et al., 2020) Image caption: (a) Shanghai, swamp type from China; (b) Toraja Spotted, swamp type from Indonesia; (c) Sylhet, swamp type from Bangladesh; (d) Mediterranean river type; (e) Murrah, river type from India; (f) Nili-Ravi, river type from Pakistan (Adopted from Zhang et al., 2020) |
Recent molecular studies have provided deeper insights into the taxonomy of water buffaloes. For instance, genetic analyses have revealed significant differences between the river and swamp buffaloes, supporting the hypothesis that they should be classified as separate species (Zhang et al., 2013). The river buffalo is now classified as Bubalus bubalis, while the swamp buffalo is classified as Bubalus kerabau. This reclassification has important implications for understanding the evolutionary history and domestication processes of these animals.
2.2 Differences between river and swamp buffalo subspecies
River and swamp buffaloes exhibit significant genetic, morphological, and ecological differences, reflecting their distinct domestication histories and adaptations to diverse environments. River buffaloes, predominantly found in the Indian subcontinent, are primarily bred for dairy production due to their high milk yield, whereas swamp buffaloes, which are widespread in Southeast Asia, are valued for their strength and endurance in labor-intensive agricultural tasks, particularly in wetland environments (Kierstein et al., 2004).
Genetic studies have provided further evidence of these distinctions. Mitochondrial DNA analyses indicate that river and swamp buffaloes belong to separate genetic clades, supporting the hypothesis of independent domestication events (Lei et al., 2007; Kumar et al., 2007). Additionally, karyotype analysis has revealed a fundamental chromosomal difference: river buffaloes possess 50 chromosomes, while swamp buffaloes have 48, reinforcing their classification as genetically distinct lineages. These genetic differences have important implications for breeding strategies and conservation efforts, as maintaining the genetic integrity of both subspecies is crucial for sustaining biodiversity and ensuring long-term population viability.
2.3 Phylogenetic placement within the bovidae family
The phylogenetic classification of water buffaloes within the Bovidae family has been a topic of extensive investigation, with molecular analyses playing a key role in elucidating their evolutionary relationships. Studies utilizing mitochondrial DNA and microsatellite markers have provided robust evidence that water buffaloes share a closer genetic affinity with sheep (Ovis aries) than with other domesticated bovids such as cattle (Bos taurus) (Arif et al., 2012).
Molecular clock analyses suggest that water buffaloes diverged from the cattle lineage approximately 21 million years ago, while their separation from the sheep lineage occurred around 0.5 million years ago. These findings refine our understanding of the evolutionary trajectory of water buffaloes, positioning them as a distinct lineage within Bovidae. The phylogenetic tree topology further underscores their unique genetic heritage, emphasizing their divergence from other domesticated species and highlighting their evolutionary significance (Lin et al., 2020).
3 Fossil Records and Evolutionary History
3.1 Overview of fossil evidence related to water buffalo evolution
Fossil evidence provides critical insights into the evolutionary history of water buffaloes, with remains found primarily in the Indian subcontinent and Southeast Asia. Early ancestors of modern water buffaloes are believed to have been widely distributed across these regions, adapting to diverse ecological niches over time (Dong et al., 2014). While traditional perspectives suggested that water buffaloes were first domesticated in Neolithic China approximately 7 000 years ago, recent zooarchaeological studies have challenged this notion. Emerging evidence indicates that domestication may have occurred in multiple regions, including South Asia and Southeast Asia, supporting a model of parallel domestication events (Yang et al., 2008).
Phylogenetic analyses of ancient DNA extracted from water buffalo remains in North China reveal that these ancient buffaloes were not direct ancestors of modern domesticated buffalo populations. Instead, they were closely related to an extinct wild species, Bubalus mephistopheles, suggesting that the domestication process involved multiple progenitor species across different geographical regions. This complex domestication history has contributed to the genetic diversity observed in contemporary water buffalo populations, influencing their adaptability and resilience.
3.2 Comparative insights from other Bovidae species fossils
Comparative fossil studies within the Bovidae family provide additional context for understanding the evolutionary origins of water buffaloes. The Bovidae family, which includes cattle, sheep, and goats, shares a common ancestral lineage with water buffaloes. Phylogenetic analyses of conserved microsatellite sequences suggest that Bubalus bubalis exhibits a closer genetic relationship with Ovis aries (sheep) than with other domestic species such as cattle and goats. This supports the hypothesis that water buffaloes diverged from a common ancestor with sheep and goats after the subfamily Bovinae, which includes cattle, had already separated from the broader Bovidae lineage.
The divergence times estimated from genetic data indicate that water buffalo separated from cattle approximately 5.8 to 9.8 million years ago (Maceachern et al., 2009; Mintoo et al., 2019). This timeline is consistent with the fossil record, which shows distinct evolutionary paths for different Bovidae species. The genetic and fossil evidence together highlight the complex evolutionary relationships within the Bovidae family and underscore the unique evolutionary trajectory of water buffalo (Iannuzzi et al., 2000).
3.3 Challenges in correlating fossil data with molecular evidence
One of the primary challenges in correlating fossil data with molecular evidence lies in the discrepancies between the two types of data. Fossil records provide physical evidence of past species and their geographical distribution, but they are often incomplete and subject to varying interpretations. In contrast, molecular evidence, such as DNA sequences, offers detailed insights into genetic relationships and evolutionary timelines but relies on the availability of well-preserved genetic material (Kikkawa et al., 1997).
For water buffalo, the fossil record suggests a broad distribution of wild ancestors across Asia, while molecular evidence points to distinct domestication events and genetic lineages. For instance, mitochondrial DNA analyses have shown that both river and swamp buffaloes may have descended from a single domestication event in the Indian subcontinent, with subsequent introgression of wild buffalo mtDNA into domestic stocks. Genetic evidence occasionally challenges or complicates interpretations derived solely from fossil data.
The genetic diversity observed in modern water buffalo populations, as revealed through analyses of microsatellite loci and mitochondrial genomes, suggests multiple domestication events and significant gene flow between populations (Vijh et al., 2008; Lin et al., 2020). These findings underscore the complexities involved in reconciling fossil evidence with molecular data, as both sources provide complementary yet sometimes conflicting perspectives on the evolutionary history of water buffalo.
4 Molecular Markers in Phylogenetic Studies
4.1 Commonly used molecular markers
Mitochondrial DNA (mtDNA) is among the most widely employed molecular markers in phylogenetic studies of water buffalo. In particular, the mitochondrial D-loop region has been extensively analyzed to elucidate phylogenetic relationships and domestication patterns. Studies examining the complete mitochondrial D-loop sequence across various buffalo breeds have provided critical insights into their evolutionary history, indicating a primary domestication event in the Indian subcontinent, followed by introgression of wild water buffalo mtDNA into domesticated populations (Kierstein et al., 2004). Similarly, comparative mitochondrial genome sequencing has effectively differentiated river and swamp buffalo, reinforcing the hypothesis of independent domestication processes (Kumar et al., 2007).
Microsatellite markers, consisting of short, highly polymorphic DNA sequences, have also been instrumental in buffalo phylogenetics. Due to their high mutation rates and codominant inheritance, microsatellites serve as powerful tools for assessing genetic diversity and population structure. Studies utilizing microsatellite loci have identified distinct genetic clusters among buffalo breeds, supporting taxonomic classifications based on morphological traits and geographic distribution (Vijh et al., 2008). Moreover, microsatellite analyses have provided insights into the genetic structure and evolutionary history of buffalo populations across different regions, revealing evidence of hybridization and distinct phylogenetic clustering patterns (Mishra et al., 2015).
4.2 Genome-wide approaches in buffalo phylogenetics
The advent of genome-wide approaches has significantly enhanced the resolution and accuracy of buffalo phylogenetic studies. Whole-genome sequencing and comparative genomics have yielded comprehensive insights into the evolutionary history and genetic variation of water buffalo. For instance, sequencing of the lowland anoa genome, alongside multiple buffalo genomes, has clarified the phylogenetic divergence between river and swamp buffalo, suggesting that these taxa separated during the Pleistocene epoch (Curaudeau et al., 2021). Additionally, genome-wide studies have revealed higher levels of heterozygosity and genetic variation than previously recognized, refining our understanding of buffalo phylogenetics.
Genome-wide association studies (GWAS) and population genomics have also provided valuable insights into trait evolution and adaptation in buffalo. A notable example is the identification of a LINE-1 transposon insertion in the ASIP gene, which has been linked to the white coat phenotype in swamp buffalo, illustrating the role of recent genetic transposition events in buffalo evolution (Liang et al., 2020). These genome-wide approaches not only advance our understanding of phylogenetic relationships but also offer crucial information for conservation and breeding programs aimed at maintaining genetic diversity and adaptability.
4.3 Comparison of molecular markers and their resolution in phylogenetic studies
Different molecular markers vary in their resolution and applicability for phylogenetic studies. Mitochondrial DNA, particularly the D-loop region, has been widely used for high-resolution analyses of maternal lineages, effectively distinguishing between river and swamp buffalo. However, mtDNA alone provides a limited perspective, as it only traces maternal inheritance and does not account for recombination or nuclear genomic variation. Consequently, integrating mtDNA with nuclear DNA markers, such as microsatellites and nuclear gene sequences, offers a more holistic approach to reconstructing buffalo phylogenetic relationships.
Microsatellite markers, owing to their high polymorphism and codominant inheritance, are particularly useful for assessing genetic diversity and population structure. They have been instrumental in identifying genetic clusters and elucidating evolutionary relationships that align with geographic and phenotypic classifications. Nevertheless, microsatellites may have limitations in resolving deeper phylogenetic relationships due to their short sequences and rapid mutation rates.
Among available methodologies, genome-wide approaches, including whole-genome sequencing and GWAS, provide the most comprehensive resolution for phylogenetic research. These techniques capture extensive genetic variation across the entire genome, enabling detailed analyses of evolutionary history, population differentiation, and trait inheritance (Liang et al., 2020). Although genome-wide approaches require greater computational and financial resources, they offer unprecedented insights into the complex phylogenetic dynamics of water buffalo, facilitating more precise evolutionary reconstructions and conservation strategies.
5 Phylogeographic Patterns and Global Dispersal
5.1 Evidence of domestication origins
The domestication of water buffalo is a subject of significant interest, with molecular evidence pointing towards South Asia as the primary center of domestication. Studies analyzing mitochondrial DNA (mtDNA) and nuclear DNA variations have provided insights into the origins of both river and swamp buffalo. For instance, research indicates that both types of buffalo likely descended from a single domestication event in the Indian subcontinent, with subsequent introgression of wild water buffalo mtDNA into domestic stocks (Kierstein et al., 2004). This is further supported by the genetic analysis of buffalo populations in north-east India, which revealed distinct clustering patterns that suggest a hybrid zone extending from north-east India to South-East Asia (Mishra et al., 2015).
Additionally, the phylogenetic relationships inferred from genomic data, including mitochondrial and Y-chromosomal sequences, highlight the distinct evolutionary paths of river and swamp buffalo. The river buffalo, primarily found in the Indian subcontinent and Mediterranean countries, and the swamp buffalo, prevalent in China and Southeast Asia, are considered to have diverged rapidly during the Pleistocene epoch (Curaudeau et al., 2021). This divergence aligns with the hypothesis that the Indian subcontinent served as a primary domestication center for river buffalo, whereas the domestication of swamp buffalo likely took place in the border regions between China and Indochina.
5.2 Routes of global dispersal for river and swamp buffalo
The global dispersal of river and swamp buffaloes has been shaped by their distinct domestication origins and subsequent human-facilitated movements. River buffalo, domesticated in the Indian subcontinent, gradually expanded westward into regions such as Egypt, the Balkans, and Italy (Zhang et al., 2020). Genetic studies indicate that river buffalo populations exhibit a relatively weak phylogeographic structure, suggesting a high degree of gene flow and phenotypic diversity resulting from extensive human-mediated dispersal.
In contrast, the dispersal of swamp buffalo followed different routes, corresponding to their domestication in the China-Indochina border region. Genetic evidence suggests that swamp buffaloes migrated southward through Peninsular Malaysia to Sumatra, Java, and Sulawesi, while simultaneously expanding northward through China and eventually reaching Taiwan, the Philippines, and Borneo (Zhang et al., 2011). This dispersal pattern is supported by the pronounced geographic differentiation observed in swamp buffalo populations, which exhibit limited gene flow and high genetic diversity across both maternal and paternal lineages. The distinct dispersal trajectories of river and swamp buffaloes underscore the influence of their domestication centers and the role of human migration and trade in shaping their global distribution.
5.3 Modern distribution patterns and genetic differentiation
The contemporary distribution of water buffaloes reflects the historical processes of domestication and dispersal, with significant genetic differentiation persisting between river and swamp buffalo populations. River buffaloes are predominantly found in the Indian subcontinent, the Mediterranean region, and parts of South America, where they were introduced for agricultural purposes (Zhang et al., 2020). These populations exhibit relatively high gene flow and phenotypic variation, as indicated by the clustering of diverse breeds in genetic studies, reflecting a weaker phylogeographic structure.
Swamp buffaloes, by contrast, are primarily distributed across China, Southeast Asia, and parts of Oceania. These populations exhibit strong genetic differentiation, with distinct maternal and paternal lineages indicative of limited gene flow between geographically separated populations (Zhang et al., 2016). The highest levels of genetic diversity within swamp buffalo populations are observed in the China-Indochina border region, supporting the hypothesis that this area represents the primary domestication center. The presence of ancestral haplotypes in these regions further reinforces their significance in the evolutionary history of swamp buffaloes.
6 Case Study Analysis: Convergent Evolution in Coat Color of Swamp Buffaloes
6.1 Research background
The domesticated water buffalo (Bubalus bubalis) represents a pivotal species in global agriculture, particularly in Asia, with its two major types: the river and swamp buffaloes (Zhang et al. 2020). Among the unique traits of swamp buffaloes is the white coat phenotype, historically attributed to cultural and religious preferences in countries like China and Indonesia. This case study explores the genetic mechanism underlying this phenotype through molecular evidence, revealing a LINE-1 transposon insertion in the ASIP gene as the causal factor and drawing parallels to similar mechanisms in cattle.
6.2 Genetic mechanism of white coat phenotype
Recent genomic and transcriptomic analyses demonstrated that the white coat phenotype in swamp buffaloes is linked to a specific LINE-1 transposon insertion in the ASIP (agouti signaling protein) gene (Figure 2) (Liang et al., 2020). This 2 809-bp-long LINE-1 insertion functions as a strong proximal promoter, resulting in a significant upregulation of ASIP expression in white buffalo skin. This overexpression inhibits melanocyte maturation, preventing melanin production, and leading to the absence of pigment in skin and hair. The LINE-1 element introduces a chimeric transcript by splicing into the ASIP coding exon, a feature distinct from black buffaloes, which lack this insertion.
|
Figure 2 Identification of the LINE-1 insertion in ASIP of white buffaloes (Adopted from Liang et al., 2020) Image caption: (A) Significantly higher expression of ASIP in white buffalo skin (W1, W2, and W3) than in black buffalo skin (B1, B2, and B3) revealed by RNA-seq and qPCR. Three experimental replicates of qPCR are shown separately (qPCR1, qPCR2, and qPCR3). (B) Distinct transcripts assembled from RNA-seq data of skin samples of white (blue) and black (red) buffaloes. (C) Full-length transcripts of ASIP generated using the RACE-PCR and characterization of the LINE-1 insertion in the white buffalo ASIP. The structure of a full-length LINE-1 element from cattle (L1-BT; Girardot et al. 2006) is shown as reference. (D) Schematic representation showing the chromosomal position of the LINE-1 insertion determined by soft-clipped reads analysis and the partial sequences of the LINE-1 insertion obtained by de novode novo assembling of the soft-clipped reads. (E) Genotyping the LINE-1 insertion using the allele-specific PCR and its perfect association with white coat phenotype. PCR products of wild allele (Normal; 296 bp) and mutant allele (Ins; 387 bp) are separated for six samples (S1−S6) using agarose gel electrophoresis (Adopted from Liang et al., 2020) |
6.3 Evidence from whole-genome sequencing
Whole-genome sequencing of swamp buffalo populations identified a genomic region on chromosome 14 strongly associated with the white coat phenotype (Liang et al., 2020). Within this region, the ASIP gene showed differential expression between white and black buffaloes, with white buffaloes exhibiting a 10-fold increase in ASIP transcription. Genotyping of the LINE-1 insertion across 91 white and 194 black buffaloes confirmed its perfect association with the white coat phenotype, establishing it as a Mendelian dominant trait.
6.4 Convergent evolution with cattle
The identification of a similar LINE-1 insertion affecting the ASIP gene in cattle provides evidence of convergent evolution in coat color across the Bovini tribe. Although the specific LINE-1 insertions in buffalo and cattle differ in terms of sequence length, genomic location, and evolutionary origin, both function as strong promoters that lead to ASIP overexpression. This suggests that transposable elements such as LINE-1 have played a significant role in regulating pigmentation genes across multiple species, shaping phenotypic diversity through evolutionary processes.
6.5 Implications for evolution and conservation
This case exemplifies the broader role of transposable elements in contributing to phenotypic variation and adaptive traits in domesticated species. The emergence of the white coat phenotype in swamp buffaloes represents a relatively recent genetic change, illustrating the dynamic nature of the domesticated animal genome. From a conservation and breeding perspective, understanding the genetic underpinnings of such traits can inform selective breeding strategies and genetic resource management, ultimately aiding in the preservation of biodiversity and the sustainable improvement of water buffalo populations.
7 Advances in Whole-Genome Sequencing and Comparative Genomics
7.1 Current progress in sequencing the water buffalo genome
Recent advances in whole-genome sequencing have significantly expanded our understanding of the Bubalus bubalis genome. The draft genome sequence of the river buffalo has been assembled, revealing a genome size of approximately 2.77 Gb, with a contig N50 of 25 kb and a scaffold N50 of 6.9 Mbp. This assembly has facilitated the annotation of 24 613 genes, providing a comprehensive foundation for future functional genomic research (Mintoo et al., 2019).
Additionally, the sequencing of the lowland anoa (Bubalus depressicornis) genome has provided comparative insights into the genetic distinctions between river and swamp buffaloes. A study incorporating three genomic datasets—including the mitochondrial genome, Y-chromosomal genes, and nuclear genes—has helped infer phylogenetic relationships while also revealing elevated levels of heterozygosity and sequence errors in the water buffalo genome (Curaudeau et al., 2021). These genomic resources have enhanced our ability to identify genetic variations and reconstruct the evolutionary history of water buffalo, while also shedding light on their relationship to other members of the Bovidae family.
7.2 Insights from comparative genomics with other domesticated species
Comparative genomic analyses have provided valuable insights into the evolutionary divergence and genetic differentiation of water buffalo in relation to other domesticated species. Phylogenetic studies indicate that water buffalo diverged from cattle (Bos taurus) approximately 5.8 to 9.8 million years ago, highlighting distinct evolutionary trajectories for these species (Mintoo et al., 2019). Furthermore, population genomic analyses of river, swamp, and hybrid buffaloes in northeastern India have revealed substantial genetic structuring, with swamp-type buffalo clustering closely with Chinese swamp buffalo, whereas river buffalo formed a distinct genetic group, illustrating a complex phylogeographic history (Mishra et al., 2015).
Comparative genomic research has also provided key insights into the genetic basis of specific phenotypic traits. For instance, the identification of a LINE-1 insertion in the ASIP gene as the genetic basis for the white coat phenotype in swamp buffalo underscores the role of transposable elements in trait evolution. Interestingly, a similar LINE-1 insertion has been detected in cattle, suggesting the presence of a convergent molecular mechanism influencing coat color variation within the Bovini tribe (Liang et al., 2020).
7.3 Implications for evolutionary adaptation and domestication traits
The integration of whole-genome sequencing and comparative genomics has profound implications for understanding the evolutionary adaptations and domestication traits of water buffalo. Phylogenetic analyses based on mitochondrial and nuclear DNA have provided strong evidence supporting the independent domestication events of river and swamp buffalo, with the former originating in the Indian subcontinent and the latter in the China-Southeast Asia region (Kierstein et al., 2004). These findings reinforce the notion that domestication occurred in multiple geographic centers, contributing to the genetic diversity observed in contemporary buffalo populations.
The identification of genetic variations associated with domestication traits, such as coat color, has direct applications for breeding and conservation programs. The discovery of the ASIP gene mutation responsible for the white coat phenotype in swamp buffalo, for example, highlights the potential of genomic selection for desirable traits in livestock breeding (Liang et al., 2020). Moreover, the pronounced genetic differentiation among geographically distinct buffalo populations underscores the importance of conservation strategies aimed at preserving unique genetic lineages and preventing genetic erosion (Zhang et al., 2011).
8 Challenges in Phylogenetic Studies of Water Buffalo
8.1 Incomplete lineage sorting and its impact on phylogenetic interpretations
Incomplete lineage sorting (ILS) poses a major challenge in the phylogenetic reconstruction of water buffalo. ILS occurs when ancestral genetic polymorphisms are randomly sorted into descendant lineages, leading to discordance between gene trees and the species tree. This phenomenon complicates efforts to accurately infer evolutionary relationships. For example, comparative genomic analyses of the lowland anoa, river buffalo, and swamp buffalo have revealed high levels of heterozygosity and sequence errors, indicative of ILS (Curaudeau et al., 2021). The presence of ILS can lead to conflicting phylogenetic signals, necessitating the use of extensive genomic datasets and sophisticated analytical approaches to disentangle genuine evolutionary relationships from artifacts of ancestral polymorphism.
Mitochondrial DNA studies have also highlighted complexities in the phylogenetic history of water buffalo, with some analyses revealing evidence of introgression and incomplete lineage sorting across different breeds (Kierstein et al., 2004). To address these challenges, researchers must integrate multiple genetic markers and employ advanced computational methods to achieve more accurate phylogenetic reconstructions.
8.2 Limitations of current molecular techniques and data availability
Despite significant advances in genomic research, limitations in molecular techniques and data availability continue to hinder phylogenetic studies of water buffalo. While microsatellite markers have provided valuable insights into genetic diversity, their limited resolution restricts their ability to resolve deeper evolutionary relationships (Vijh et al., 2008). Similarly, while the draft genome assembly of river buffalo represents a crucial resource, its fragmented nature (as indicated by contig and scaffold N50 values) poses challenges for the precise identification of genomic regions relevant to phylogenetics (Mintoo et al., 2019).
To overcome these limitations, there is a need for more extensive genomic datasets and the application of newer sequencing technologies, such as long-read sequencing and pangenomics, to enhance the resolution of buffalo phylogenetics.
8.3 Issues related to sample bias and geographical coverage
Sample bias and insufficient geographical representation pose additional challenges in water buffalo phylogenetic research. Many genetic studies have focused on specific breeds or populations, resulting in an incomplete picture of the species’ overall genetic diversity. For example, mitochondrial DNA analyses of buffalo populations in Brazil and Italy have provided valuable insights but lack broader representation from key domestication centers (Zhang et al., 2020). Similarly, studies focusing on Indian buffalo populations have revealed genetic structuring but may not fully capture the diversity of swamp buffalo populations in Southeast Asia (Mishra et al., 2015).
To address these gaps, future research should prioritize comprehensive sampling across diverse geographic regions, ensuring that underrepresented populations are included. Standardizing sampling methods and increasing international collaboration will also improve the robustness of phylogenetic analyses and enhance our understanding of water buffalo evolution.
9 Concluding Remarks and Future Directions
Molecular studies have played a pivotal role in advancing our understanding of the phylogenetic evolution of water buffalo (Bubalus bubalis), shedding light on distinct evolutionary trajectories and domestication events. Analyses of mitochondrial DNA (mtDNA) and nuclear markers have provided compelling evidence for the divergence of river and swamp buffaloes, with domestication likely occurring in separate centers—the Indian subcontinent for river buffalo and the China-Southeast Asia region for swamp buffalo. Furthermore, genome-wide studies have refined our understanding of genetic relationships within and between these populations, revealing hybridization zones and distinct genetic clusters. These findings underscore the complexity of water buffalo phylogeny and highlight its significant implications for genetic diversity, domestication history, and conservation strategies.
To develop a more comprehensive understanding of water buffalo evolution, future research should integrate molecular findings with archaeological and ecological evidence. Archaeological discoveries, including ancient remains and settlement patterns, can provide valuable context for molecular data, offering insights into domestication timelines and historical migration routes. Likewise, ecological studies can help elucidate the influence of environmental pressures on genetic diversity and adaptive traits. By combining these interdisciplinary approaches, researchers can construct a more nuanced and holistic evolutionary framework, addressing existing knowledge gaps and reconciling conflicting interpretations between molecular and fossil evidence.
The intersection of conservation genomics and evolutionary biology presents promising opportunities for advancing our understanding of water buffalo genetics. Emerging sequencing technologies and comparative genomic analyses can be leveraged to identify key genetic markers associated with desirable traits, such as disease resistance and environmental adaptability. These insights can inform targeted breeding programs aimed at enhancing genetic resilience and sustainability, particularly for genetically unique or endangered populations. Additionally, further exploration of hybridization dynamics and their role in shaping genetic diversity could provide crucial information for managing and preserving genetic resources. Given the historical and ecological significance of water buffalo, future studies should prioritize the inclusion of underrepresented geographical regions to ensure a more complete and accurate representation of their genetic variation and evolutionary history.
![]() Acknowledgments
Acknowledgments
The authors sincerely thank their colleague Anita W.W. from the research team for the assistance provided in the collection of literature and materials for this study.
Conflict of Interest Disclosure
The authors affirm that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Arif I., Bakir M., and Khan H., 2012, Inferring the phylogeny of bovidae using mitochondrial dna sequences: resolving power of individual genes relative to complete genomes, Evolutionary Bioinformatics Online, 8: 139-150.
https://doi.org/10.4137/EBO.S8897
PMid:22399841 PMCid:PMC3290115
Curaudeau M., Rozzi R., and Hassanin A., 2021, The genome of the lowland anoa (Bubalus depressicornis) illuminates the origin of river and swamp buffalo, Molecular Phylogenetics and Evolution, 107170.
https://doi.org/10.1016/j.ympev.2021.107170
PMid:33798669
Dong W., Liu J., Zhang L., and Xu Q., 2014, The early pleistocene water buffalo associated with Gigantopithecus from Chongzuo in southern China. Quaternary International, 354: 86-93.
https://doi.org/10.1016/j.quaint.2013.12.054
Girardot M., Guibert S., Laforet M.P., Gallard Y., Larroque H., and Oulmouden A., 2006, The insertion of a full-length Bos taurus LINE element is responsible for a transcriptional deregulation of the Normande Agouti gene, Pigment Cell Res. 19(4): 346-355.
https://doi.org/10.1111/j.1600-0749.2006.00312.x
PMid:16827753
Iannuzzi L., Meo G., Perucatti A., Incarnato D., Schibler L., and Cribiu E., 2000 , Comparative FISH mapping of bovid X chromosomes reveals homologies and divergences between the subfamilies Bovinae and Caprinae, Cytogenetic and Genome Research, 89:171-176.
https://doi.org/10.1159/000015607
PMid:10965117
Kierstein G., Vallinoto M., Silva A., Schneider M., Iannuzzi L., and Brenig B., 2004, Analysis of mitochondrial D-loop region casts new light on domestic water buffalo (Bubalus bubalis) phylogeny, Molecular Phylogenetics and Evolution, 30(2): 308-324.
https://doi.org/10.1016/S1055-7903(03)00221-5
PMid:14715223
Kikkawa Y., Yonekawa H., Suzuki H., and Amano T., 1997, Analysis of genetic diversity of domestic water buffaloes and anoas based on variations in the mitochondrial gene for cytochrome b, Animal Genetics, 28: 195-201.
https://doi.org/10.1111/j.1365-2052.1997.00101.x
Kumar S., Nagarajan M., Sandhu J., Kumar N., Behl V., and Nishanth G., 2007, Mitochondrial DNA analyses of Indian water buffalo support a distinct genetic origin of river and swamp buffalo, Animal Genetics, 38(3): 227-232.
https://doi.org/10.1111/j.1365-2052.2007.01602.x
PMid:17459014
Lei C., Lei C., Zhang W., Chen H., Lu F., Liu R., Yang X., Zhang H., Liu Z., Yao L., Lu Z., and Zhao Z., 2007, Independent maternal origin of Chinese swamp buffalo (Bubalus bubalis), Animal Genetics, 38(2): 97-102.
https://doi.org/10.1111/j.1365-2052.2007.01567.x
PMid:17313588
Liang D., Zhao P., Si J., Fang L., Pairo-Castineira E., Hu X., Xu Q., Hou Y., Gong Y., Liang Z., Tian B., Mao H., Yindee M., Faruque M., Kongvongxay S., Khamphoumee S., Liu G., Wu D., Barker J., Han J., and Zhang Y., 2020, Genomic analysis revealed a convergent evolution of line-1 in coat color: a case study in water buffaloes (Bubalus bubalis), Molecular Biology and Evolution, 38: 1122-1136.
https://doi.org/10.1093/molbev/msaa279
PMid:33212507 PMCid:PMC7947781
Lin Q., Li M., Wang Y., and Qiu H., 2020, Determination and phylogenetic analysis of the complete mitochondrial genome of Bubalus bubalis Linnaeus, 1758 breed Murrah (Artiodactyla: Bovidae), Mitochondrial DNA, Part B, Resources, 5: 432-433.
https://doi.org/10.1080/23802359.2019.1704181
PMid:33426275 PMCid:PMC7755322
Maceachern S., McEwan J., and Goddard M., 2009, Phylogenetic reconstruction and the identification of ancient polymorphism in the Bovini tribe (Bovidae, Bovinae), BMC Genomics, 10: 177-177.
https://doi.org/10.1186/1471-2164-10-177
PMid:19393045 PMCid:PMC2694835
Mintoo A., Zhang H., Chen C., Moniruzzaman M., Deng T., Anam M., Huque Q., Guang X., Wang P., Zhong Z., Han P., Khatun A., Awal T., Gao Q., and Liang X., 2019, Draft genome of the river water buffalo, Ecology and Evolution, 9: 3378-3388.
https://doi.org/10.1002/ece3.4965
PMid:30962899 PMCid:PMC6434576
Mishra B., Dubey P., Prakash B., Kathiravan P., Goyal S., Sadana D., Das G., Goswami R., Bhasin V., Joshi B., and Kataria R., 2015, Genetic analysis of river, swamp and hybrid buffaloes of north-east India throw new light on phylogeography of water buffalo (Bubalus bubalis), Journal of Animal Breeding and Genetics = Zeitschrift fur Tierzuchtung und Zuchtungsbiologie, 132(6), 454-466.
https://doi.org/10.1111/jbg.12141
PMid:25780854
Nagarajan M., Nimisha K., and Kumar S., 2015, Mitochondrial DNA Variability of domestic river buffalo (Bubalus bubalis) populations: genetic evidence for domestication of river buffalo in indian subcontinent, Genome Biology and Evolution, 7: 1252-1259.
https://doi.org/10.1093/gbe/evv067
PMid:25900921 PMCid:PMC4453062
Sikdar S., Das T., Sajib E.H., Rahman K.M.U., Siddiki A.Z., and Uddin M.B., 2021, Multi-OMICS and molecular biology perspective in Buffalo genome, J. Buffalo Sci, 10: 21-31.
https://doi.org/10.6000/1927-520X.2021.10.04
Vijh R.K., Tantia M.S., Mishra B., and Bharani Kumar S.T., 2008, Genetic relationship and diversity analysis of Indian water buffalo (Bubalus bubalis), Journal of Animal Science, 86(7): 1495-1502.
https://doi.org/10.2527/jas.2007-0321
PMid:18344309
Yang D., Liu L., Chen X., and Speller C., 2008, Wild or domesticated: DNA analysis of ancient water buffalo remains from north China, Journal of Archaeological Science, 35: 2778-2785.
https://doi.org/10.1016/j.jas.2008.05.010
Yi Z., Barker J., Vankan D., and Yuan Z., 2013, The swamp buffalo: domestication, dispersal, and genetic differentiation, Buffalo Bulletin, 32: 671-674.
Zhang Y., Colli L., and Barker J.S.F., 2020, Asian water buffalo: domestication, history and genetics. Animal genetics, 51(2): 177-191.
https://doi.org/10.1111/age.12911
PMid:31967365
Zhang Y., Lu Y., Yindee M., Li K., Kuo H., Ju Y., Ye S., Faruque M., Li Q., Wang Y., Cuong V., Pham L., Bouahom B., Yang B., Liang X., Cai Z., Vankan D., Manatchaiworakul W., Kowlim N., Duangchantrasiri S., Wajjwalku W., Colenbrander B., Zhang Y., Beerli P., Lenstra J., and Barker J., 2016, Strong and stable geographic differentiation of swamp buffalo maternal and paternal lineages indicates domestication in the China/Indochina border region, Molecular Ecology, 25(7): 1530-1550.
https://doi.org/10.1111/mec.13518
PMid:26677084
Zhang Y., Vankan D., Zhang Y., and Barker J., 2011, Genetic differentiation of water buffalo (Bubalus bubalis) populations in China, Nepal and south-east Asia: inferences on the region of domestication of the swamp buffalo, Animal Genetics, 42(4): 366-377.
https://doi.org/10.1111/j.1365-2052.2010.02166.x
PMid:21749419
.png)
. HTML
Associated material
. Readers' comments
Other articles by authors
. Xiaoli Chen
. Shiqiang Huang
Related articles
. Water buffalo ( Bubalus bubalis )
. Phylogeny
. Molecular evidence
. Domestication and hybridization
. Genetic diversity
Tools
. Post a comment
.png)


